
Wave Life Sciences Corporate Presentation November 5, 2019 Exhibit 99.2

Forward-looking statements This document contains forward-looking statements. All statements other than statements of historical facts contained in this document, including statements regarding possible or assumed future results of operations, preclinical and clinical studies, business strategies, research and development plans, collaborations and partnerships, regulatory activities and timing thereof, competitive position, potential growth opportunities, use of proceeds and the effects of competition are forward-looking statements. These statements involve known and unknown risks, uncertainties and other important factors that may cause the actual results, performance or achievements of Wave Life Sciences Ltd. (the “Company”) to be materially different from any future results, performance or achievements expressed or implied by the forward-looking statements. In some cases, you can identify forward-looking statements by terms such as “may,” “will,” “should,” “expect,” “plan,” “aim,” “anticipate,” “could,” “intend,” “target,” “project,” “contemplate,” “believe,” “estimate,” “predict,” “potential” or “continue” or the negative of these terms or other similar expressions. The forward-looking statements in this presentation are only predictions. The Company has based these forward-looking statements largely on its current expectations and projections about future events and financial trends that it believes may affect the Company’s business, financial condition and results of operations. These forward-looking statements speak only as of the date of this presentation and are subject to a number of risks, uncertainties and assumptions, including those listed under Risk Factors in the Company’s Form 10-K and other filings with the SEC, some of which cannot be predicted or quantified and some of which are beyond the Company’s control. The events and circumstances reflected in the Company’s forward-looking statements may not be achieved or occur, and actual results could differ materially from those projected in the forward-looking statements. Moreover, the Company operates in a dynamic industry and economy. New risk factors and uncertainties may emerge from time to time, and it is not possible for management to predict all risk factors and uncertainties that the Company may face. Except as required by applicable law, the Company does not plan to publicly update or revise any forward-looking statements contained herein, whether as a result of any new information, future events, changed circumstances or otherwise.

DESIGN & OPTIMIZE SEQUENCE STEREOCHEMISTRY CHEMISTRY Targeting genetically defined diseases with stereopure oligonucleotides Building fully integrated genetic medicines company led by neurology development programs Lead clinical program: Suvodirsen Phase 2/3 trial ongoing for DMD (exon 51); program on development path toward US and global approvals Advancing additional exon skipping candidates for DMD Commercialization activities underway Lead clinical program: Two Phase 1b/2a trials ongoing for Huntington’s disease using differentiated allele-selective approach Advancing C9orf72 candidate for ALS and FTD SNP3 (HD) and ATXN3 (SCA3) Advancing USH2A exon-skipping program for Usher Syndrome Stereopure oligonucleotides across multiple therapeutic modalities Silencing | Splicing | Editing Neuromuscular CNS Ophthalmology 100% global rights Takeda 50:50 option 100% global rights
Through iterative analysis of in vitro and in vivo outcomes and artificial intelligence-driven predictive modeling, Wave continues to define design principles that are deployed across programs to rapidly develop and manufacture clinical candidates that meet pre-defined product profiles DESIGN Unique ability to construct stereopure oligonucleotides with one defined and consistent profile Enables Wave to target genetically defined diseases with stereopure oligonucleotides across multiple therapeutic modalities OPTIMIZE A deep understanding of how the interplay among oligonucleotide sequence, chemistry, and backbone stereochemistry impacts key pharmacological properties SEQUENCE STEREOCHEMISTRY CHEMISTRY
WAVE RATIONAL DESIGN Control of stereochemistry enables the design and manufacture of oligonucleotides with one defined and consistent profile Designing the optimal, stereopure medicine STANDARD OLIGONUCLEOTIDE APPROACHES Pharmacologic properties include >500,000 permutations in every dose Impact: Unreliable therapeutic effects Unintended off-target effects Impact: Potential for best-in-class medicines that can address difficult-to-treat diseases
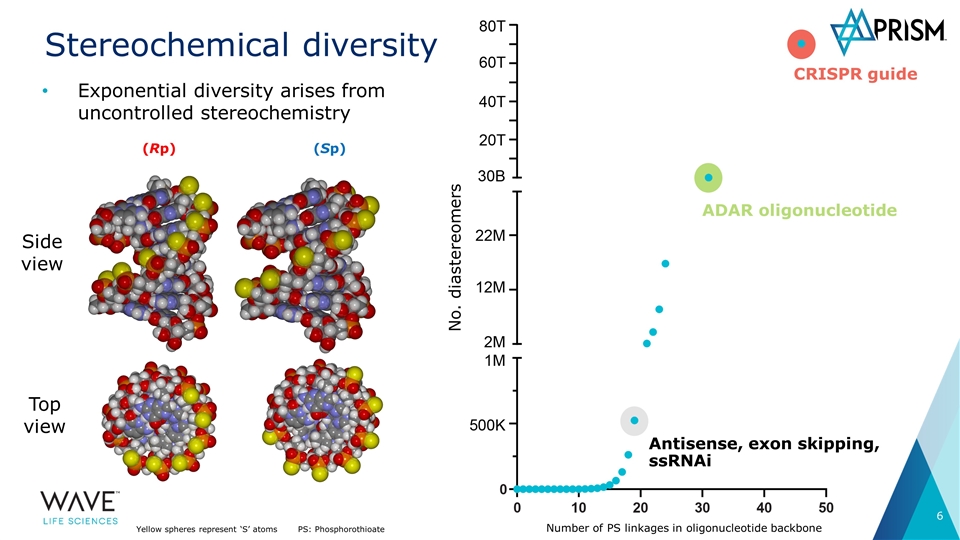
Stereochemical diversity Number of PS linkages in oligonucleotide backbone Antisense, exon skipping, ssRNAi CRISPR guide ADAR oligonucleotide No. diastereomers Exponential diversity arises from uncontrolled stereochemistry (Rp) (Sp) Top view Side view Yellow spheres represent ‘S’ atomsPS: Phosphorothioate
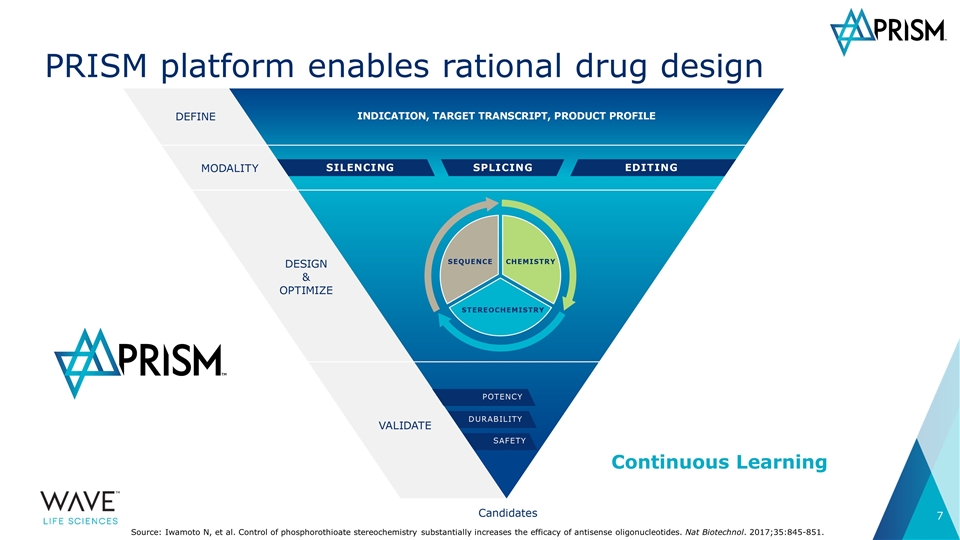
Continuous Learning PRISM platform enables rational drug design Source: Iwamoto N, et al. Control of phosphorothioate stereochemistry substantially increases the efficacy of antisense oligonucleotides. Nat Biotechnol. 2017;35:845-851.
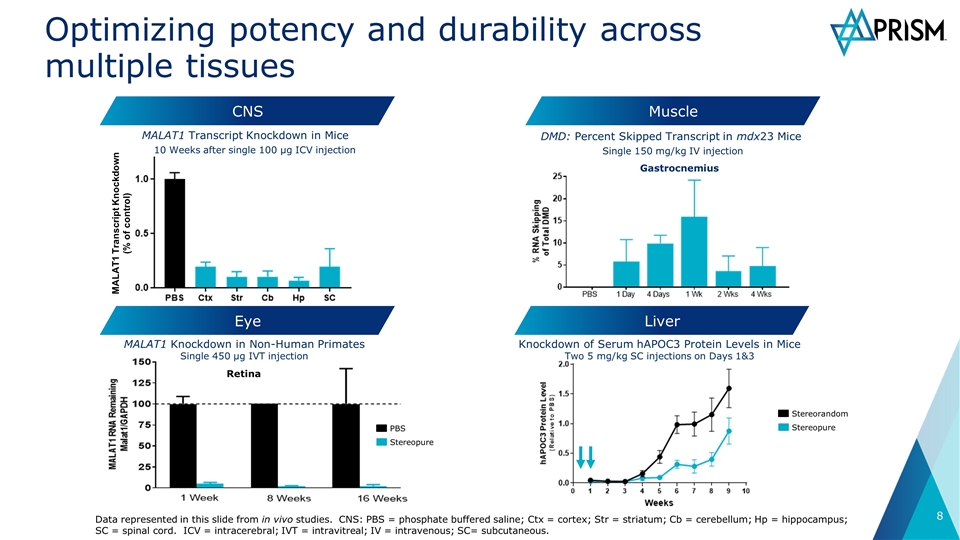
CNS Muscle Liver MALAT1 Transcript Knockdown in Mice Knockdown of Serum hAPOC3 Protein Levels in Mice Two 5 mg/kg SC injections on Days 1&3 PBS Stereopure Eye MALAT1 Knockdown in Non-Human Primates Single 450 µg IVT injection 10 Weeks after single 100 µg ICV injection DMD: Percent Skipped Transcript in mdx23 Mice Stereorandom Stereopure Single 150 mg/kg IV injection Data represented in this slide from in vivo studies. CNS: PBS = phosphate buffered saline; Ctx = cortex; Str = striatum; Cb = cerebellum; Hp = hippocampus; SC = spinal cord. ICV = intracerebral; IVT = intravitreal; IV = intravenous; SC= subcutaneous. Retina Gastrocnemius MALAT1 Transcript Knockdown (% of control) Optimizing potency and durability across multiple tissues
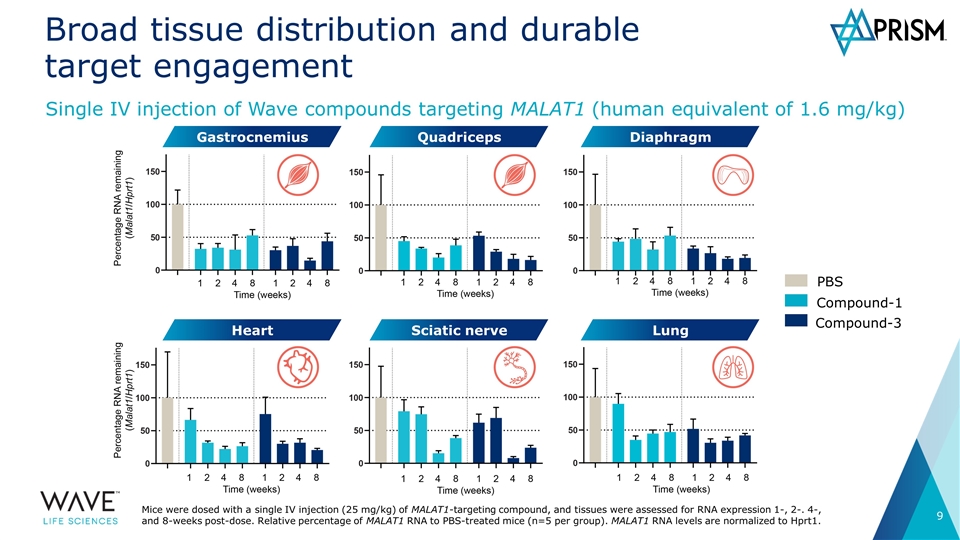
Broad tissue distribution and durable target engagement Single IV injection of Wave compounds targeting MALAT1 (human equivalent of 1.6 mg/kg) PBS Compound-1 Compound-3 Gastrocnemius Quadriceps Diaphragm Sciatic nerve Lung Heart Mice were dosed with a single IV injection (25 mg/kg) of MALAT1-targeting compound, and tissues were assessed for RNA expression 1-, 2-. 4-, and 8-weeks post-dose. Relative percentage of MALAT1 RNA to PBS-treated mice (n=5 per group). MALAT1 RNA levels are normalized to Hprt1.
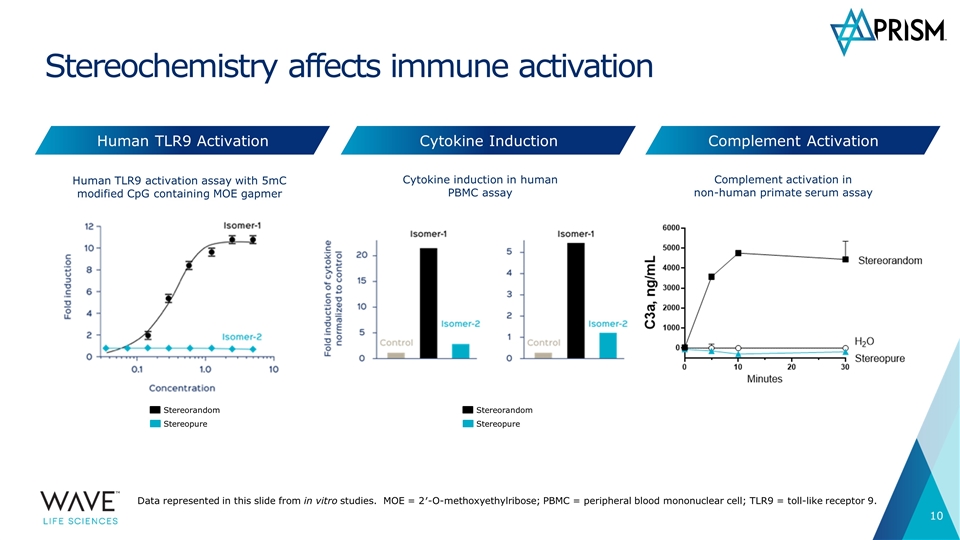
Stereochemistry allows for Human TLR9 activation assay with 5mC modified CpG containing MOE gapmer Cytokine induction in human PBMC assay Stereochemistry affects immune activation Complement Activation Human TLR9 Activation Cytokine Induction Complement activation in non-human primate serum assay Data represented in this slide from in vitro studies. MOE = 2′-O-methoxyethylribose; PBMC = peripheral blood mononuclear cell; TLR9 = toll-like receptor 9. Stereorandom Stereopure Stereorandom Stereopure
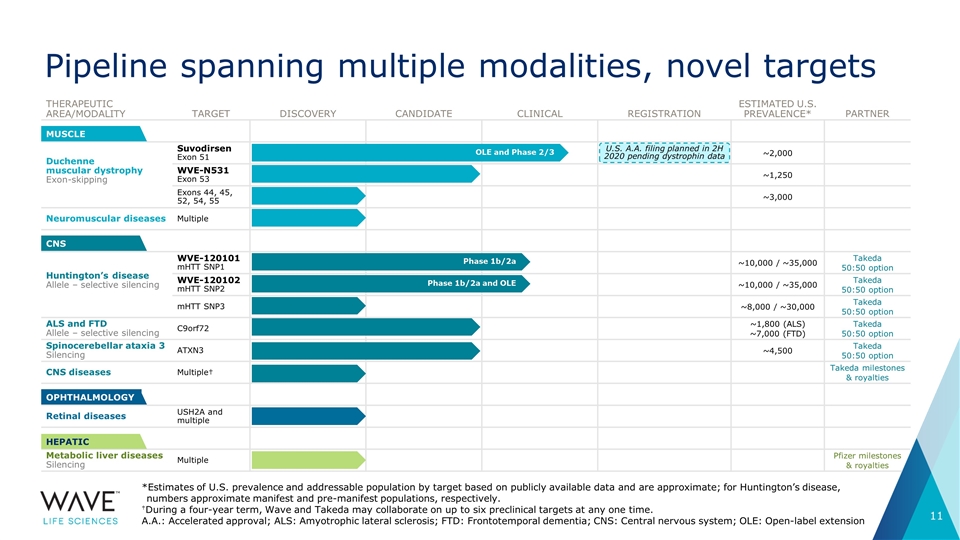
THERAPEUTIC AREA/MODALITY TARGET DISCOVERY CANDIDATE CLINICAL REGISTRATION ESTIMATED U.S. PREVALENCE* PARTNER Duchenne muscular dystrophy Exon-skipping Suvodirsen Exon 51 ~2,000 WVE-N531 Exon 53 ~1,250 Exons 44, 45, 52, 54, 55 ~3,000 Neuromuscular diseases Multiple Huntington’s disease Allele – selective silencing WVE-120101 mHTT SNP1 ~10,000 / ~35,000 Takeda 50:50 option WVE-120102 mHTT SNP2 ~10,000 / ~35,000 Takeda 50:50 option mHTT SNP3 ~8,000 / ~30,000 Takeda 50:50 option ALS and FTD Allele – selective silencing C9orf72 ~1,800 (ALS) ~7,000 (FTD) Takeda 50:50 option Spinocerebellar ataxia 3 Silencing ATXN3 ~4,500 Takeda 50:50 option CNS diseases Multiple† Takeda milestones & royalties Retinal diseases USH2A and multiple Metabolic liver diseases Silencing Multiple Pfizer milestones & royalties MUSCLE CNS OPHTHALMOLOGY HEPATIC U.S. A.A. filing planned in 2H 2020 pending dystrophin data OLE and Phase 2/3 Phase 1b/2a Phase 1b/2a and OLE Pipeline spanning multiple modalities, novel targets *Estimates of U.S. prevalence and addressable population by target based on publicly available data and are approximate; for Huntington’s disease, numbers approximate manifest and pre-manifest populations, respectively. †During a four-year term, Wave and Takeda may collaborate on up to six preclinical targets at any one time. A.A.: Accelerated approval; ALS: Amyotrophic lateral sclerosis; FTD: Frontotemporal dementia; CNS: Central nervous system; OLE: Open-label extension

Suvodirsen Duchenne Muscular Dystrophy (DMD)

DMD: a progressive, fatal childhood disorder Fatal, X-linked genetic neuromuscular disorder characterized by progressive, irreversible loss of muscle function, including heart and lung Genetic mutation in dystrophin gene prevents the production of dystrophin protein, a critical component of healthy muscle function Symptom onset in early childhood; one of the most serious genetic diseases in children worldwide Current disease modifying treatments have demonstrated minimal dystrophin expression and clinical benefit has not been established Impacts 1 in every 5,000 newborn boys each year; 20,000 new cases annually worldwide Neuro DMD Source: Parent Project Muscular Dystrophy. About Duchenne & Becker muscular dystrophy. Available at: https://www.parentprojectmd.org/care/for-healthcare-providers/. Accessed: November 2, 2018.
Exon skipping Potential benefits of stereopure oligonucleotide approach to treating Duchenne muscular dystrophy Neuro DMD Sources: Arnett ALH, et al. Mol Ther Methods Clin Dev. 2014;1:14038. doi:10.1038/mtm.2014.38. Counsell JR, et al. Sci Rep. 2017;7:79. doi: 10.1038/s41598-017-00152-5. Duan D. Mol Ther. 2018;25:2337-2356. Martinsen B, Dreyer P. Open Nurs Jrnl. 2016;10:131-138. Stitelman DH, et al. Mol Ther Methods Clin Dev. 2014;1:14040. doi:10.1038/mtm.2014.40; Aartsma-Rus A et al. J Med Genet. 2016;53(3):145-151. Scalable manufacturing Scalable manufacturing process to meet clinical and commercial supply requirements Cost of goods consistent with conventional oligonucleotide therapies Repeat administration Repeat administration may better address muscle cell turnover and need for broad distribution Functional dystrophin Production of meaningful levels of functional dystrophin protein Expected to result in therapeutic benefit Delivery Entry into cells (including progenitor cells) via free-uptake Enhanced nuclear uptake Broad tissue distribution Wave stereopure oligonucleotide RNA transcript Functional protein Dystrophin: Largest gene in human genome
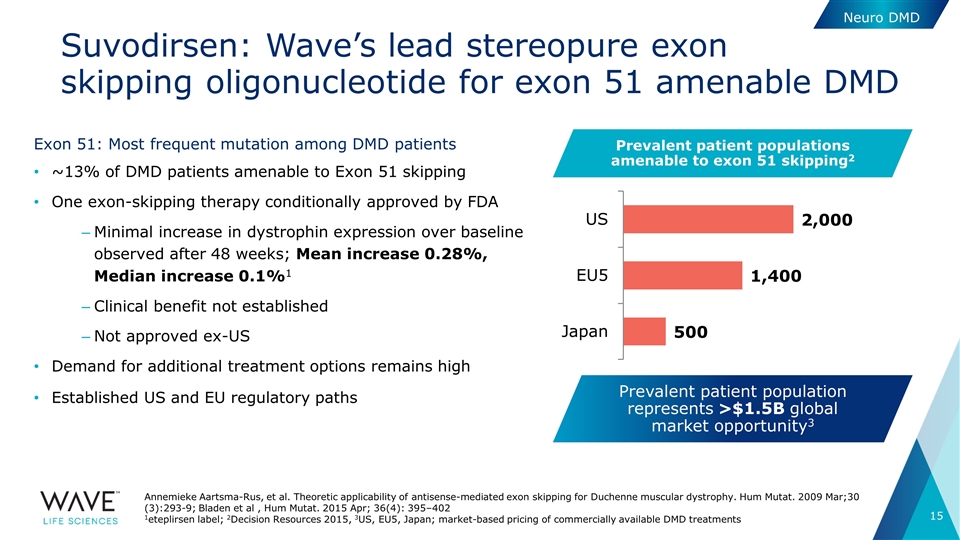
Exon 51: Most frequent mutation among DMD patients ~13% of DMD patients amenable to Exon 51 skipping One exon-skipping therapy conditionally approved by FDA Minimal increase in dystrophin expression over baseline observed after 48 weeks; Mean increase 0.28%, Median increase 0.1%1 Clinical benefit not established Not approved ex-US Demand for additional treatment options remains high Established US and EU regulatory paths Neuro DMD Annemieke Aartsma-Rus, et al. Theoretic applicability of antisense-mediated exon skipping for Duchenne muscular dystrophy. Hum Mutat. 2009 Mar;30 (3):293-9; Bladen et al , Hum Mutat. 2015 Apr; 36(4): 395–402 1eteplirsen label; 2Decision Resources 2015, 3US, EU5, Japan; market-based pricing of commercially available DMD treatments Prevalent patient population represents >$1.5B global market opportunity3 Prevalent patient populations amenable to exon 51 skipping2 Suvodirsen: Wave’s lead stereopure exon skipping oligonucleotide for exon 51 amenable DMD
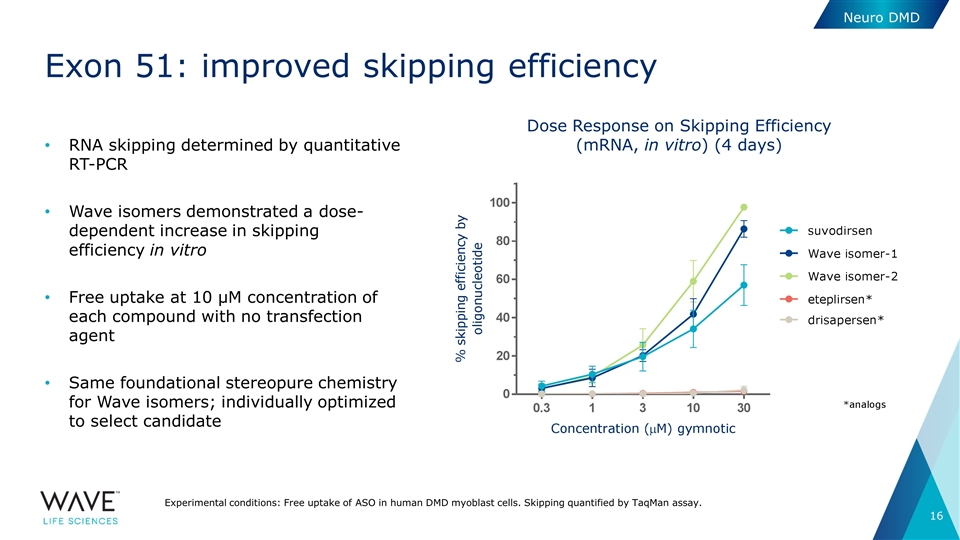
Exon 51: improved skipping efficiency RNA skipping determined by quantitative RT-PCR Wave isomers demonstrated a dose-dependent increase in skipping efficiency in vitro Free uptake at 10 µM concentration of each compound with no transfection agent Same foundational stereopure chemistry for Wave isomers; individually optimized to select candidate Neuro DMD Dose Response on Skipping Efficiency (mRNA, in vitro) (4 days) Experimental conditions: Free uptake of ASO in human DMD myoblast cells. Skipping quantified by TaqMan assay.
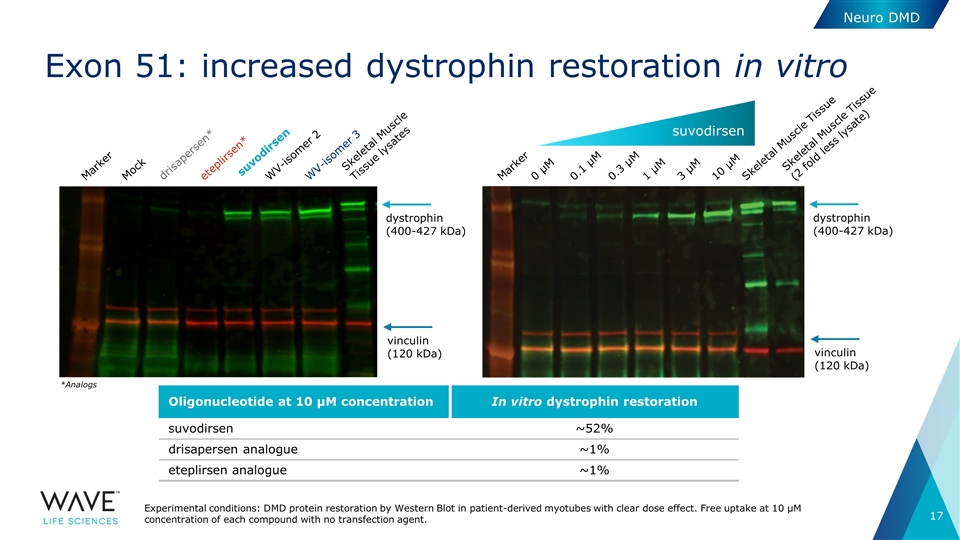

Exon 51: increased dystrophin restoration in vitro *Analogs dystrophin (400-427 kDa) vinculin (120 kDa) Marker Mock drisapersen* eteplirsen* suvodirsen WV-isomer 2 WV-isomer 3 Skeletal Muscle Tissue lysates Marker 0 µM Skeletal Muscle Tissue (2 fold less lysate) 0.1 µM 0.3 µM 1 µM 3 µM 10 µM Skeletal Muscle Tissue dystrophin (400-427 kDa) vinculin (120 kDa) Experimental conditions: DMD protein restoration by Western Blot in patient-derived myotubes with clear dose effect. Free uptake at 10 µM concentration of each compound with no transfection agent. suvodirsen Neuro DMD Oligonucleotide at 10 µM concentration In vitro dystrophin restoration suvodirsen ~52% drisapersen analogue ~1% eteplirsen analogue ~1%
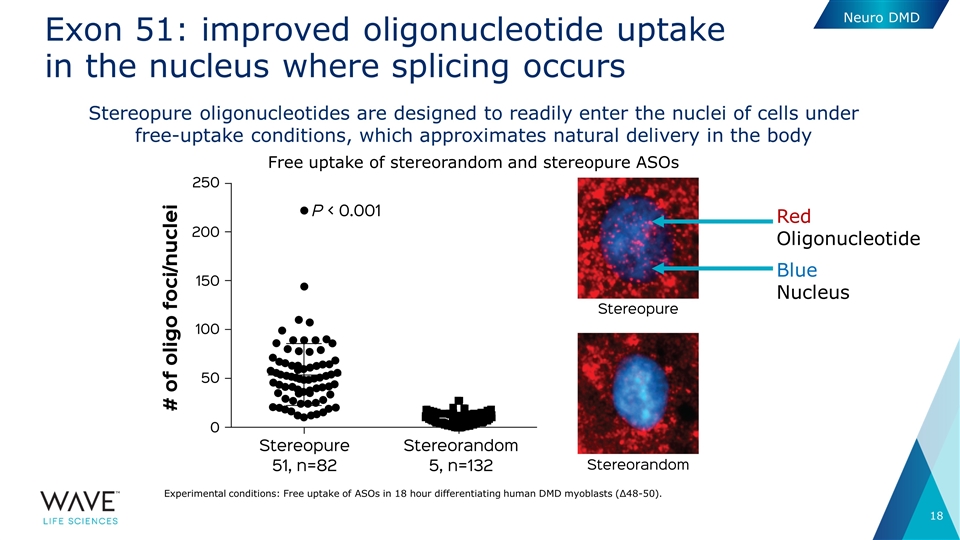

Exon 51: improved oligonucleotide uptake in the nucleus where splicing occurs Stereopure oligonucleotides are designed to readily enter the nuclei of cells under free-uptake conditions, which approximates natural delivery in the body Free uptake of stereorandom and stereopure ASOs Experimental conditions: Free uptake of ASOs in 18 hour differentiating human DMD myoblasts (Δ48-50). Red Oligonucleotide Blue Nucleus Neuro DMD
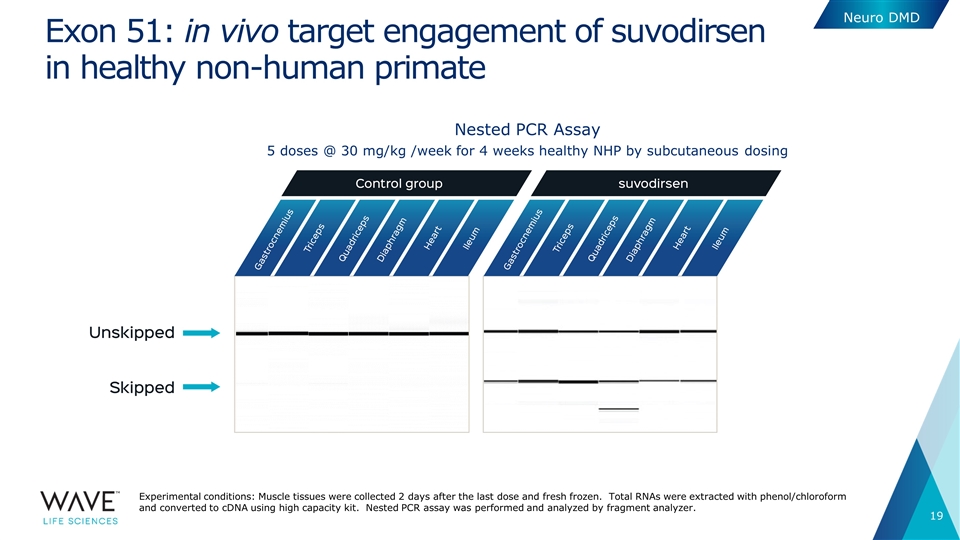
Exon 51: in vivo target engagement of suvodirsen in healthy non-human primate 5 doses @ 30 mg/kg /week for 4 weeks healthy NHP by subcutaneous dosing Nested PCR Assay Neuro DMD Experimental conditions: Muscle tissues were collected 2 days after the last dose and fresh frozen. Total RNAs were extracted with phenol/chloroform and converted to cDNA using high capacity kit. Nested PCR assay was performed and analyzed by fragment analyzer.
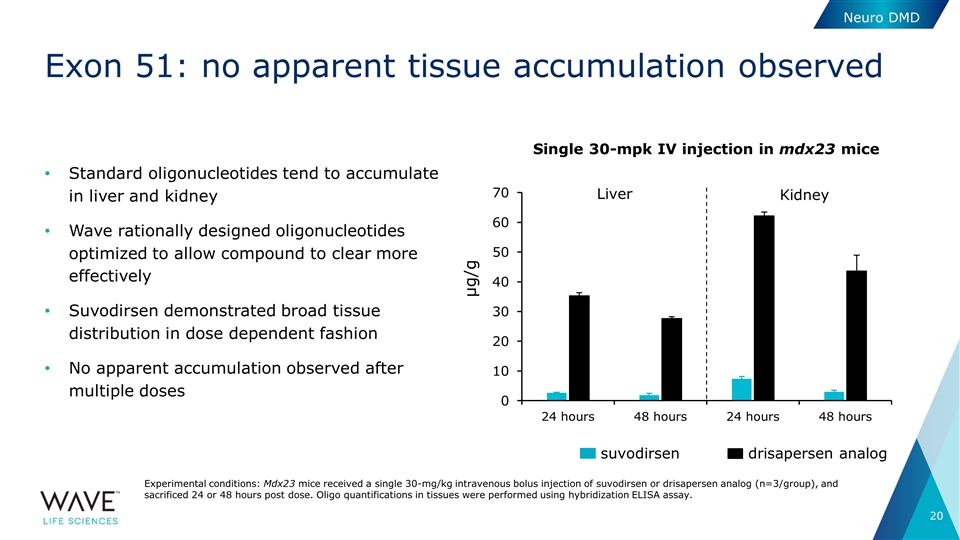
Exon 51: no apparent tissue accumulation observed Standard oligonucleotides tend to accumulate in liver and kidney Wave rationally designed oligonucleotides optimized to allow compound to clear more effectively Suvodirsen demonstrated broad tissue distribution in dose dependent fashion No apparent accumulation observed after multiple doses Neuro DMD Experimental conditions: Mdx23 mice received a single 30-mg/kg intravenous bolus injection of suvodirsen or drisapersen analog (n=3/group), and sacrificed 24 or 48 hours post dose. Oligo quantifications in tissues were performed using hybridization ELISA assay. Single 30-mpk IV injection in mdx23 mice suvodirsen drisapersen analog µg/g
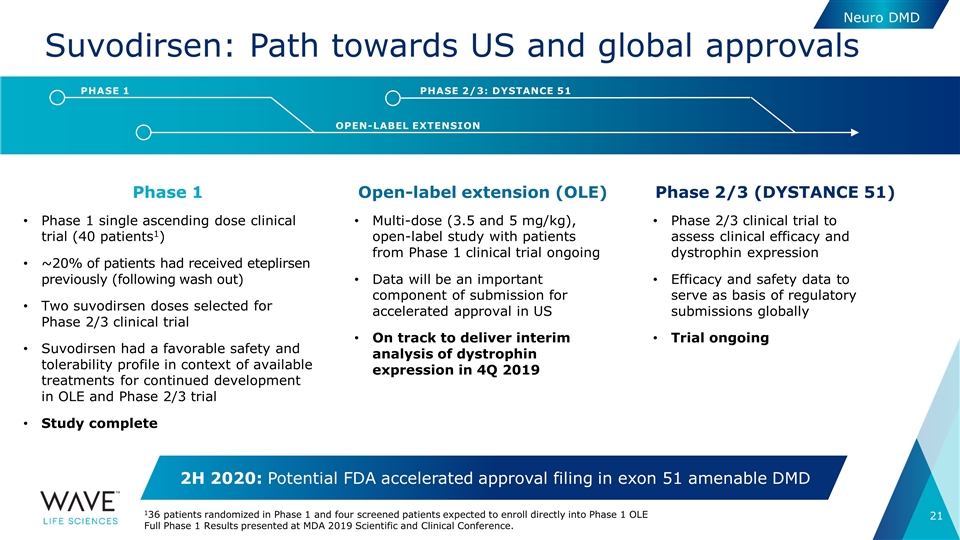

Suvodirsen: Path towards US and global approvals Phase 1 Phase 1 single ascending dose clinical trial (40 patients1) ~20% of patients had received eteplirsen previously (following wash out) Two suvodirsen doses selected for Phase 2/3 clinical trial Suvodirsen had a favorable safety and tolerability profile in context of available treatments for continued development in OLE and Phase 2/3 trial Study complete Phase 2/3 (DYSTANCE 51) Phase 2/3 clinical trial to assess clinical efficacy and dystrophin expression Efficacy and safety data to serve as basis of regulatory submissions globally Trial ongoing OPEN-LABEL EXTENSION PHASE 1 PHASE 2/3: DYSTANCE 51 Open-label extension (OLE) Multi-dose (3.5 and 5 mg/kg), open-label study with patients from Phase 1 clinical trial ongoing Data will be an important component of submission for accelerated approval in US On track to deliver interim analysis of dystrophin expression in 4Q 2019 Neuro DMD 2H 2020: Potential FDA accelerated approval filing in exon 51 amenable DMD 136 patients randomized in Phase 1 and four screened patients expected to enroll directly into Phase 1 OLE Full Phase 1 Results presented at MDA 2019 Scientific and Clinical Conference.
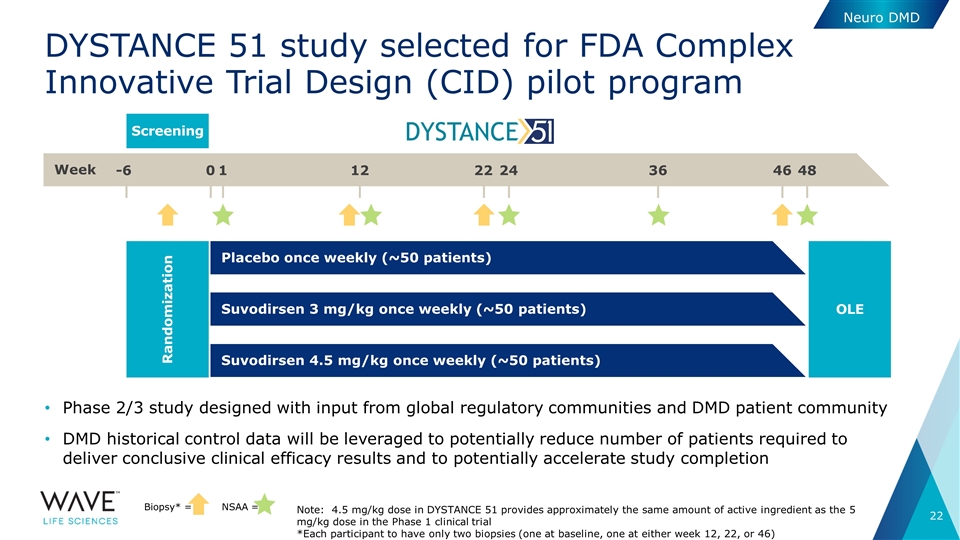
DYSTANCE 51 study selected for FDA Complex Innovative Trial Design (CID) pilot program Phase 2/3 study designed with input from global regulatory communities and DMD patient community DMD historical control data will be leveraged to potentially reduce number of patients required to deliver conclusive clinical efficacy results and to potentially accelerate study completion Neuro DMD Week -6 0 1 12 22 24 36 46 48 Screening Biopsy* = NSAA = Randomization Placebo once weekly (~50 patients) OLE Suvodirsen 3 mg/kg once weekly (~50 patients) Suvodirsen 4.5 mg/kg once weekly (~50 patients) Note: 4.5 mg/kg dose in DYSTANCE 51 provides approximately the same amount of active ingredient as the 5 mg/kg dose in the Phase 1 clinical trial *Each participant to have only two biopsies (one at baseline, one at either week 12, 22, or 46)
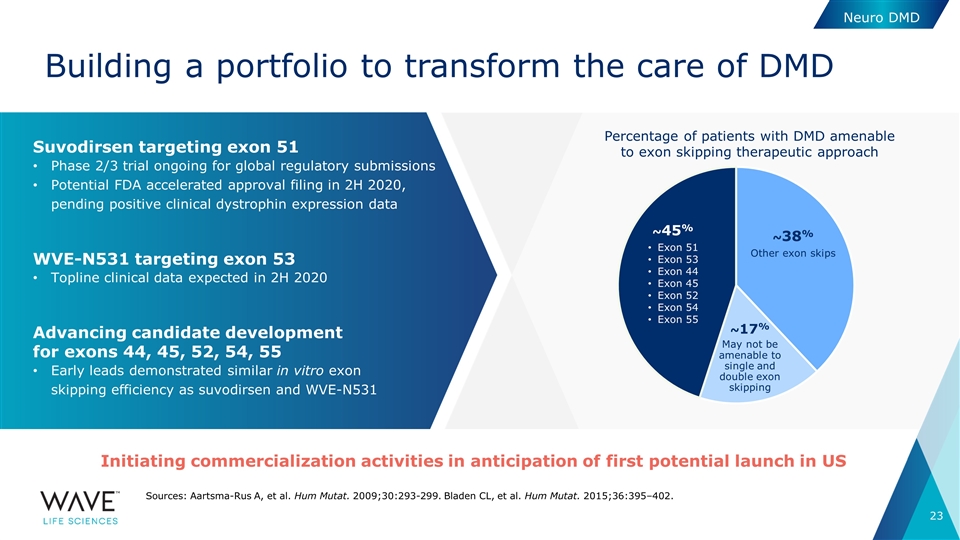
Building a portfolio to transform the care of DMD Neuro DMD Sources: Aartsma-Rus A, et al. Hum Mutat. 2009;30:293-299. Bladen CL, et al. Hum Mutat. 2015;36:395–402. Suvodirsen targeting exon 51 Phase 2/3 trial ongoing for global regulatory submissions Potential FDA accelerated approval filing in 2H 2020, pending positive clinical dystrophin expression data WVE-N531 targeting exon 53 Topline clinical data expected in 2H 2020 Advancing candidate development for exons 44, 45, 52, 54, 55 Early leads demonstrated similar in vitro exon skipping efficiency as suvodirsen and WVE-N531 Percentage of patients with DMD amenable to exon skipping therapeutic approach ~45% Exon 51 Exon 53 Exon 44 Exon 45 Exon 52 Exon 54 Exon 55 ~17% May not be amenable to single and double exon skipping ~38% Other exon skips Initiating commercialization activities in anticipation of first potential launch in US
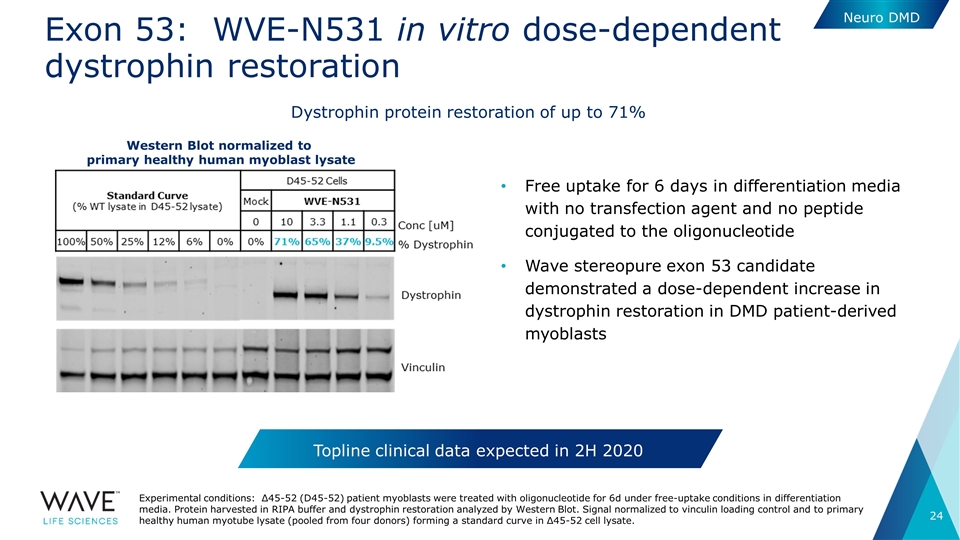

Exon 53: WVE-N531 in vitro dose-dependent dystrophin restoration Free uptake for 6 days in differentiation media with no transfection agent and no peptide conjugated to the oligonucleotide Wave stereopure exon 53 candidate demonstrated a dose-dependent increase in dystrophin restoration in DMD patient-derived myoblasts Experimental conditions: Δ45-52 (D45-52) patient myoblasts were treated with oligonucleotide for 6d under free-uptake conditions in differentiation media. Protein harvested in RIPA buffer and dystrophin restoration analyzed by Western Blot. Signal normalized to vinculin loading control and to primary healthy human myotube lysate (pooled from four donors) forming a standard curve in Δ45-52 cell lysate. Neuro DMD Topline clinical data expected in 2H 2020 Dystrophin protein restoration of up to 71% Western Blot normalized to primary healthy human myoblast lysate
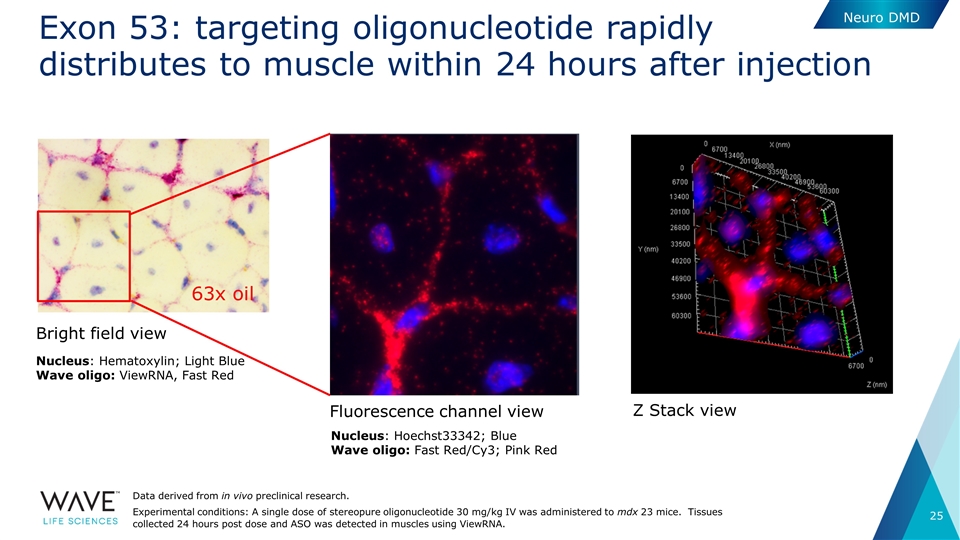
Exon 53: targeting oligonucleotide rapidly distributes to muscle within 24 hours after injection Bright field view 63x oil Nucleus: Hematoxylin; Light Blue Wave oligo: ViewRNA, Fast Red Nucleus: Hoechst33342; Blue Wave oligo: Fast Red/Cy3; Pink Red Fluorescence channel view Z Stack view Data derived from in vivo preclinical research. Experimental conditions: A single dose of stereopure oligonucleotide 30 mg/kg IV was administered to mdx 23 mice. Tissues collected 24 hours post dose and ASO was detected in muscles using ViewRNA. Neuro DMD
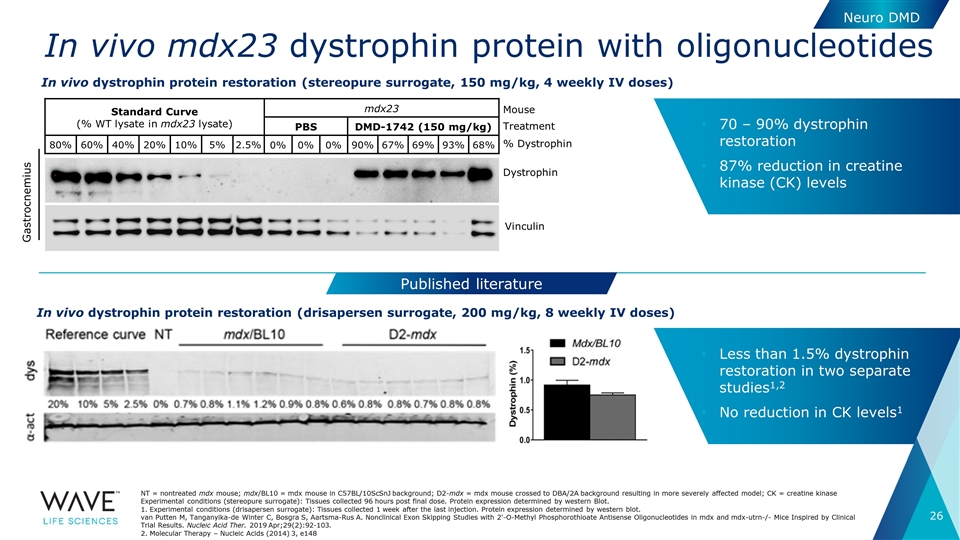
In vivo mdx23 dystrophin protein with oligonucleotides NT = nontreated mdx mouse; mdx/BL10 = mdx mouse in C57BL/10ScSnJ background; D2-mdx = mdx mouse crossed to DBA/2A background resulting in more severely affected model; CK = creatine kinase Experimental conditions (stereopure surrogate): Tissues collected 96 hours post final dose. Protein expression determined by western Blot. 1. Experimental conditions (drisapersen surrogate): Tissues collected 1 week after the last injection. Protein expression determined by western blot. van Putten M, Tanganyika-de Winter C, Bosgra S, Aartsma-Rus A. Nonclinical Exon Skipping Studies with 2'-O-Methyl Phosphorothioate Antisense Oligonucleotides in mdx and mdx-utrn-/- Mice Inspired by Clinical Trial Results. Nucleic Acid Ther. 2019 Apr;29(2):92-103. 2. Molecular Therapy – Nucleic Acids (2014) 3, e148 Gastrocnemius In vivo dystrophin protein restoration (stereopure surrogate, 150 mg/kg, 4 weekly IV doses) Standard Curve (% WT lysate in mdx23 lysate) mdx23 PBS DMD-1742 (150 mg/kg) 80% 60% 40% 20% 10% 5% 2.5% 0% 0% 0% 90% 67% 69% 93% 68% Dystrophin Vinculin % Dystrophin Treatment Mouse 70 – 90% dystrophin restoration 87% reduction in creatine kinase (CK) levels In vivo dystrophin protein restoration (drisapersen surrogate, 200 mg/kg, 8 weekly IV doses) Published literature Less than 1.5% dystrophin restoration in two separate studies1,2 No reduction in CK levels1 Neuro DMD
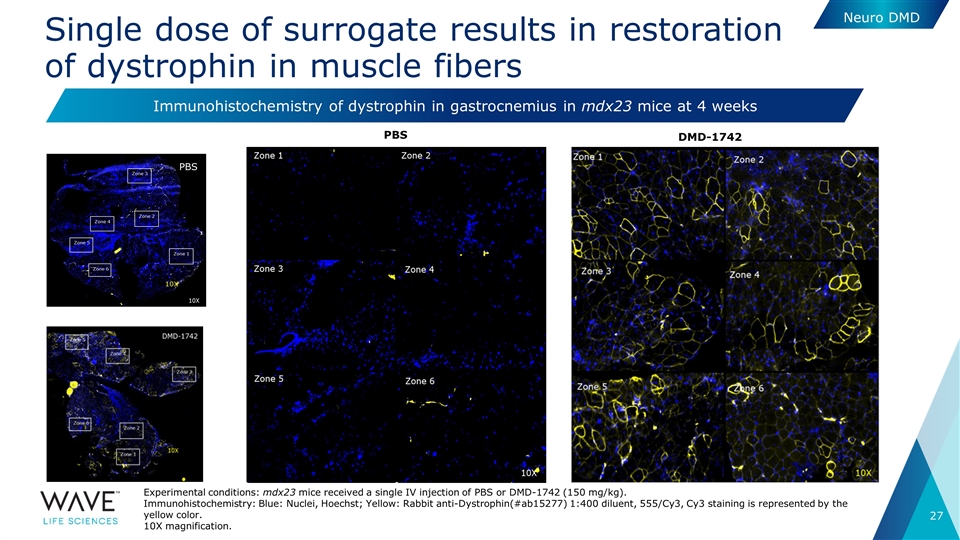
Single dose of surrogate results in restoration of dystrophin in muscle fibers Neuro DMD PBS DMD-1742 Immunohistochemistry of dystrophin in gastrocnemius in mdx23 mice at 4 weeks 10X Experimental conditions: mdx23 mice received a single IV injection of PBS or DMD-1742 (150 mg/kg). Immunohistochemistry: Blue: Nuclei, Hoechst; Yellow: Rabbit anti-Dystrophin(#ab15277) 1:400 diluent, 555/Cy3, Cy3 staining is represented by the yellow color. 10X magnification. PBS
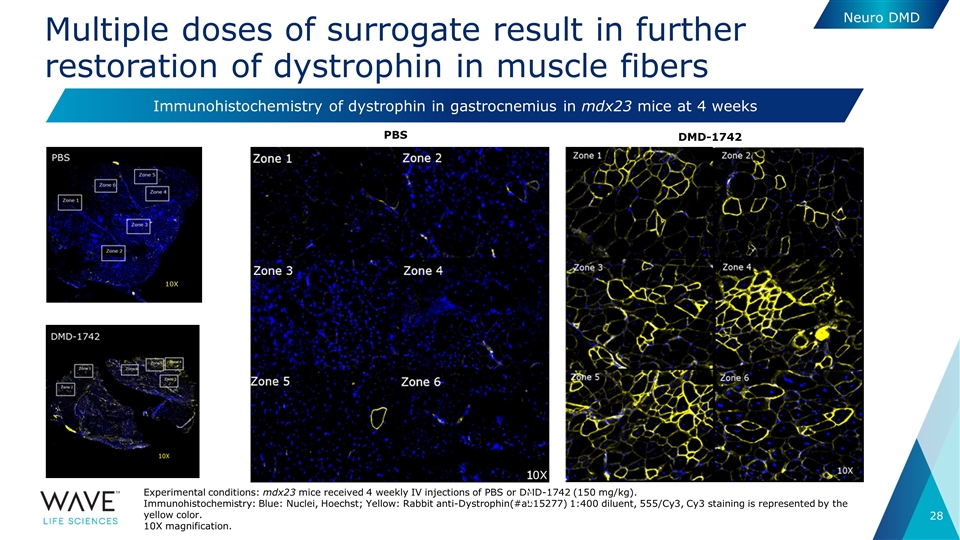
Multiple doses of surrogate result in further restoration of dystrophin in muscle fibers Experimental conditions: mdx23 mice received 4 weekly IV injections of PBS or DMD-1742 (150 mg/kg). Immunohistochemistry: Blue: Nuclei, Hoechst; Yellow: Rabbit anti-Dystrophin(#ab15277) 1:400 diluent, 555/Cy3, Cy3 staining is represented by the yellow color. 10X magnification. Neuro DMD PBS DMD-1742 Immunohistochemistry of dystrophin in gastrocnemius in mdx23 mice at 4 weeks 10X 0X

WVE-120101 WVE-120102 Huntington’s Disease
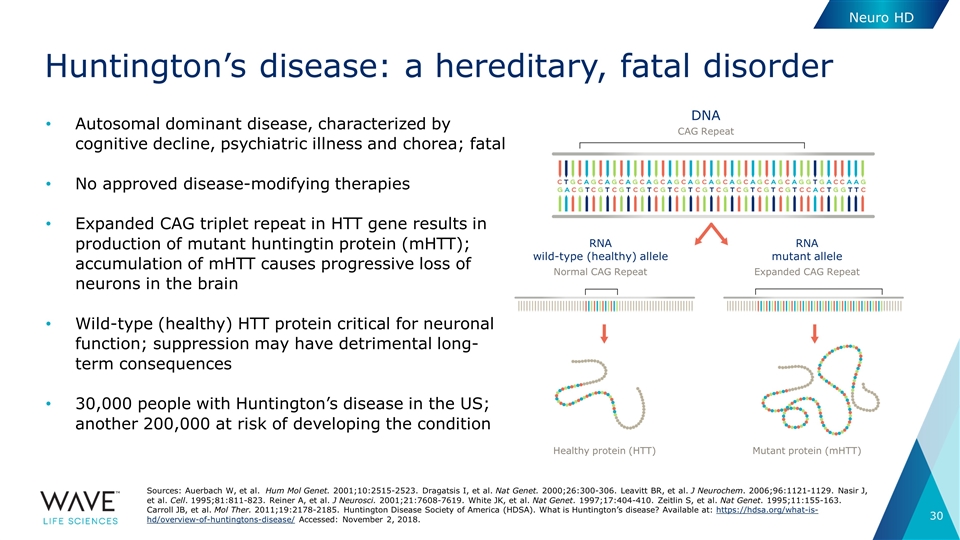
Huntington’s disease: a hereditary, fatal disorder Sources: Auerbach W, et al. Hum Mol Genet. 2001;10:2515-2523. Dragatsis I, et al. Nat Genet. 2000;26:300-306. Leavitt BR, et al. J Neurochem. 2006;96:1121-1129. Nasir J, et al. Cell. 1995;81:811-823. Reiner A, et al. J Neurosci. 2001;21:7608-7619. White JK, et al. Nat Genet. 1997;17:404-410. Zeitlin S, et al. Nat Genet. 1995;11:155-163. Carroll JB, et al. Mol Ther. 2011;19:2178-2185. Huntington Disease Society of America (HDSA). What is Huntington’s disease? Available at: https://hdsa.org/what-is-hd/overview-of-huntingtons-disease/ Accessed: November 2, 2018. DNA CAG Repeat RNA wild-type (healthy) allele RNA mutant allele Normal CAG Repeat Expanded CAG Repeat Healthy protein (HTT) Mutant protein (mHTT) Neuro HD Autosomal dominant disease, characterized by cognitive decline, psychiatric illness and chorea; fatal No approved disease-modifying therapies Expanded CAG triplet repeat in HTT gene results in production of mutant huntingtin protein (mHTT); accumulation of mHTT causes progressive loss of neurons in the brain Wild-type (healthy) HTT protein critical for neuronal function; suppression may have detrimental long-term consequences 30,000 people with Huntington’s disease in the US; another 200,000 at risk of developing the condition

Utilize association between single nucleotide polymorphisms (SNPs) and genetic mutations to specifically target errors in genetic disorders, including Huntington’s disease (HD) Potential to provide treatment for up to 80% of HD population Wave approach: novel, allele-selective silencing Source: Kay, et al. Personalized gene silencing therapeutics for Huntington disease. Clin Genet. 2014;86:29–36. Neuro HD Aims to lower mHTT transcript while leaving healthy wild-type HTT relatively intact Allele-selectivity possible by targeting SNPs associated with expanded long CAG repeat in HTT gene RNase H and ASO:RNA RNA mutant allele

Two simultaneous Phase 1b/2a clinical trials Neuro HD PRECISION-HD is a global clinical program consisting of the PRECISION-HD1 trial evaluating WVE-120101 targeting SNP1 and the PRECISION-HD2 trial evaluating WVE-120102 targeting SNP2 Two parallel, multicenter, double-blind, randomized, placebo-controlled Phase 1b/2a clinical trials for WVE-120101 and WVE-120102, administered intrathecally, with single-ascending dose and multiple-ascending dose portions Primary objective: Assess safety and tolerability of intrathecal doses in early manifest HD patients Key additional objectives: Measurement of total HTT and mHTT; exploratory pharmacokinetic (PK), pharmacodynamic (PD), clinical and MRI endpoints Key inclusion criteria: age ≥25 to ≤65, stage I or II HD who have screened positively for the presence of SNP1 or SNP2 Expected to enroll approximately 50 patients per trial Topline data expected to include: summary of clinical safety results, degree of mHTT protein lowering in CSF at 20 weeks, the ratio of total HTT versus mHTT in CSF at 20 weeks Open-label extension (OLE) study initiated for PRECISION-HD2 outside of the U.S. to allow for continued dosing and clinical assessments PRECISION-HD1 OLE expected to initiate in 2020 Intend to explore efficacy in early manifest and pre-manifest HD patient populations Topline data readout for PRECISION-HD2 expected by YE 2019
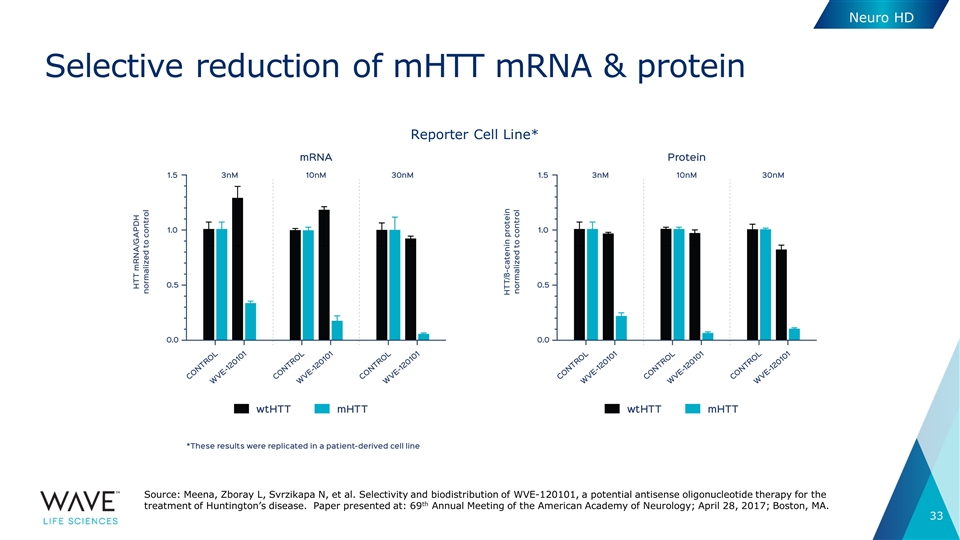
Selective reduction of mHTT mRNA & protein Reporter Cell Line* Neuro HD Source: Meena, Zboray L, Svrzikapa N, et al. Selectivity and biodistribution of WVE-120101, a potential antisense oligonucleotide therapy for the treatment of Huntington’s disease. Paper presented at: 69th Annual Meeting of the American Academy of Neurology; April 28, 2017; Boston, MA.
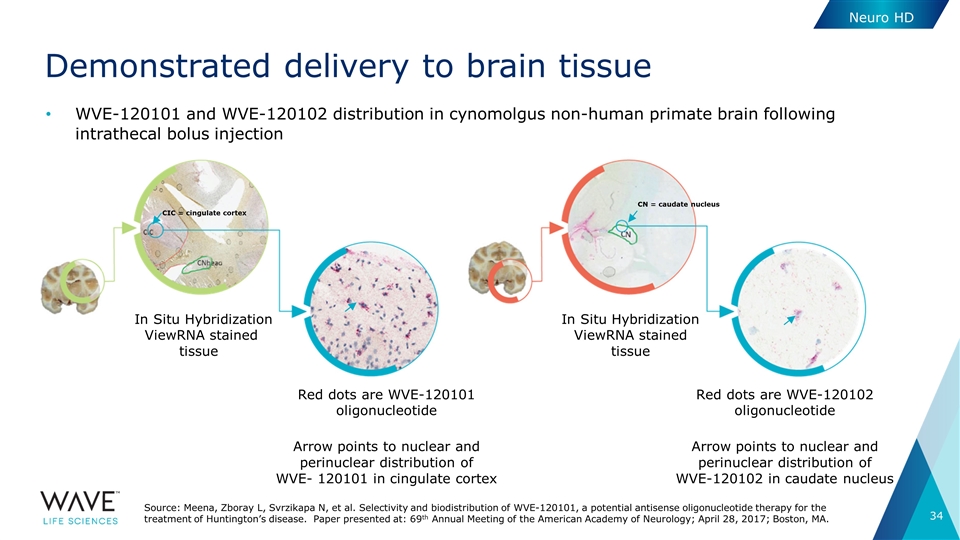
Demonstrated delivery to brain tissue WVE-120101 and WVE-120102 distribution in cynomolgus non-human primate brain following intrathecal bolus injection In Situ Hybridization ViewRNA stained tissue Red dots are WVE-120102 oligonucleotide Arrow points to nuclear and perinuclear distribution of WVE-120102 in caudate nucleus Red dots are WVE-120101 oligonucleotide Arrow points to nuclear and perinuclear distribution of WVE- 120101 in cingulate cortex CIC = cingulate cortex In Situ Hybridization ViewRNA stained tissue Neuro HD CN = caudate nucleus Source: Meena, Zboray L, Svrzikapa N, et al. Selectivity and biodistribution of WVE-120101, a potential antisense oligonucleotide therapy for the treatment of Huntington’s disease. Paper presented at: 69th Annual Meeting of the American Academy of Neurology; April 28, 2017; Boston, MA.

SNP3 Preclinical Program Huntington’s Disease
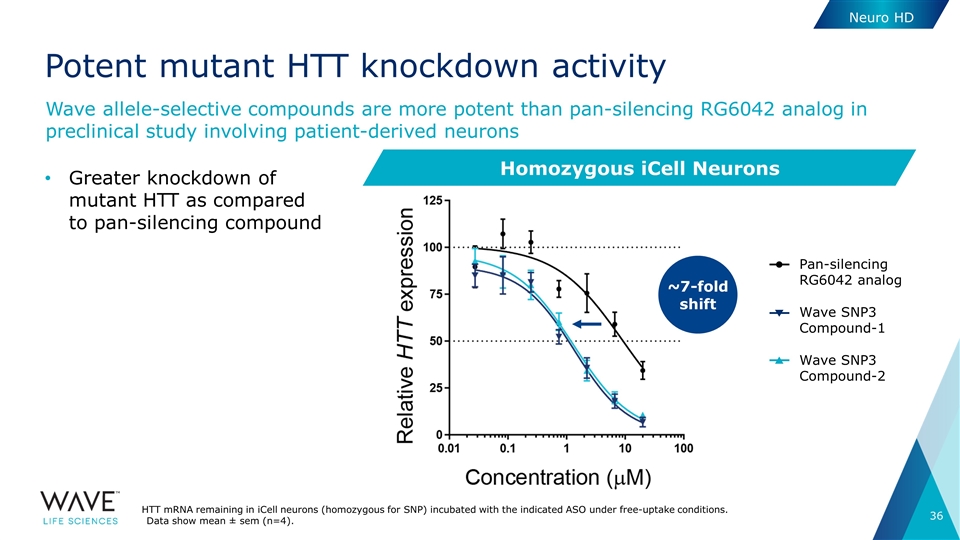
Potent mutant HTT knockdown activity Greater knockdown of mutant HTT as compared to pan-silencing compound Wave allele-selective compounds are more potent than pan-silencing RG6042 analog in preclinical study involving patient-derived neurons ~7-fold shift Pan-silencing RG6042 analog Wave SNP3 Compound-1 Wave SNP3 Compound-2 HTT mRNA remaining in iCell neurons (homozygous for SNP) incubated with the indicated ASO under free-uptake conditions. Data show mean ± sem (n=4). Homozygous iCell Neurons Neuro HD
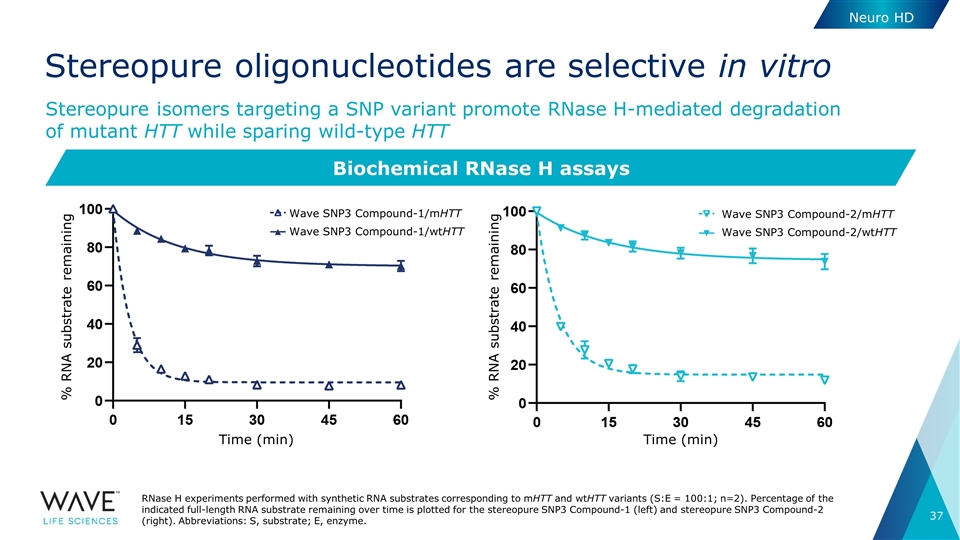
Stereopure oligonucleotides are selective in vitro Stereopure isomers targeting a SNP variant promote RNase H-mediated degradation of mutant HTT while sparing wild-type HTT Biochemical RNase H assays RNase H experiments performed with synthetic RNA substrates corresponding to mHTT and wtHTT variants (S:E = 100:1; n=2). Percentage of the indicated full-length RNA substrate remaining over time is plotted for the stereopure SNP3 Compound-1 (left) and stereopure SNP3 Compound-2 (right). Abbreviations: S, substrate; E, enzyme. Wave SNP3 Compound-1/mHTT Wave SNP3 Compound-1/wtHTT Time (min) % RNA substrate remaining Wave SNP3 Compound-2/mHTT Wave SNP3 Compound-2/wtHTT % RNA substrate remaining Time (min) Neuro HD
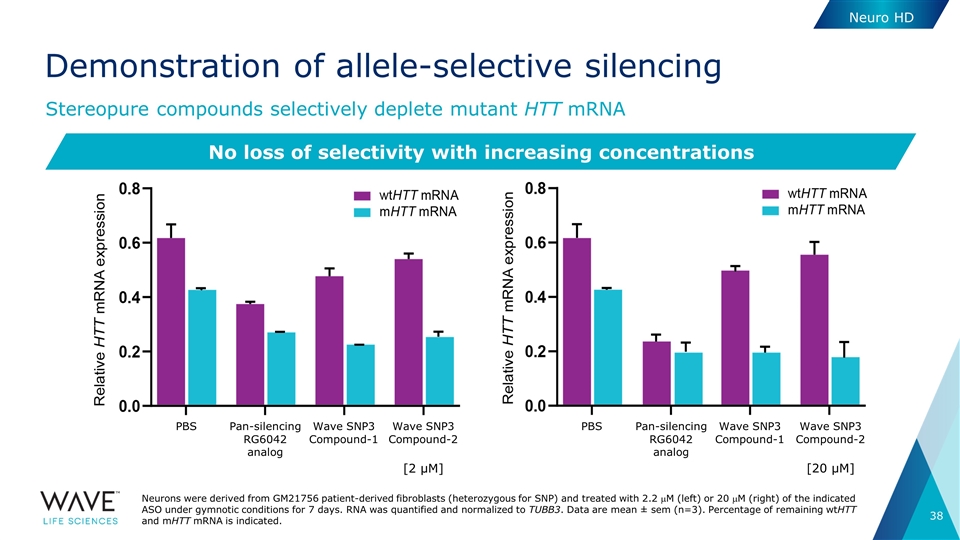
Demonstration of allele-selective silencing Stereopure compounds selectively deplete mutant HTT mRNA Neurons were derived from GM21756 patient-derived fibroblasts (heterozygous for SNP) and treated with 2.2 mM (left) or 20 mM (right) of the indicated ASO under gymnotic conditions for 7 days. RNA was quantified and normalized to TUBB3. Data are mean ± sem (n=3). Percentage of remaining wtHTT and mHTT mRNA is indicated. No loss of selectivity with increasing concentrations [20 µM] PBS PBS Pan-silencing RG6042 analog Pan-silencing RG6042 analog Wave SNP3 Compound-1 Wave SNP3 Compound-2 Wave SNP3 Compound-1 Wave SNP3 Compound-2 [2 µM] Neuro HD
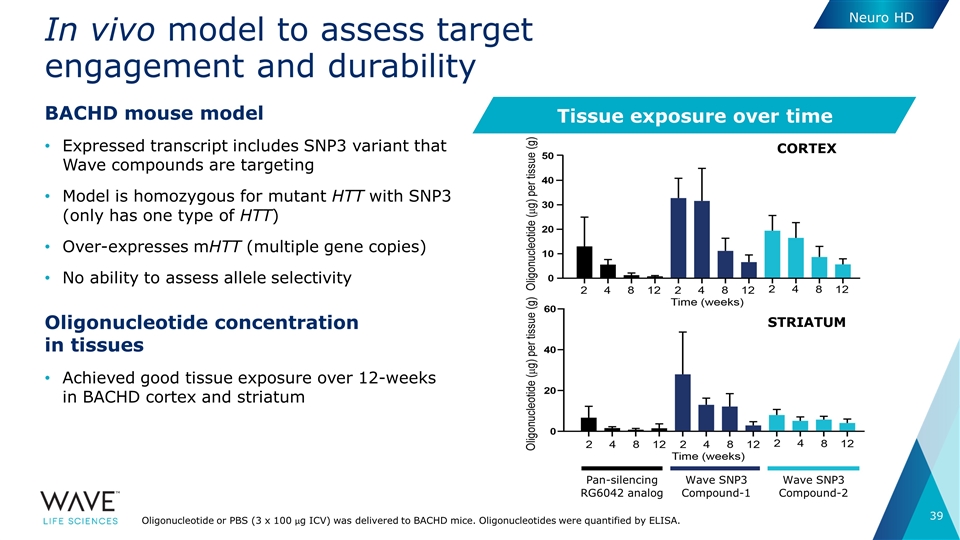
In vivo model to assess target engagement and durability BACHD mouse model Expressed transcript includes SNP3 variant that Wave compounds are targeting Model is homozygous for mutant HTT with SNP3 (only has one type of HTT) Over-expresses mHTT (multiple gene copies) No ability to assess allele selectivity Oligonucleotide concentration in tissues Achieved good tissue exposure over 12-weeks in BACHD cortex and striatum Oligonucleotide or PBS (3 x 100 µg ICV) was delivered to BACHD mice. Oligonucleotides were quantified by ELISA. Tissue exposure over time CORTEX STRIATUM Pan-silencing RG6042 analog Wave SNP3 Compound-1 Wave SNP3 Compound-2 Neuro HD
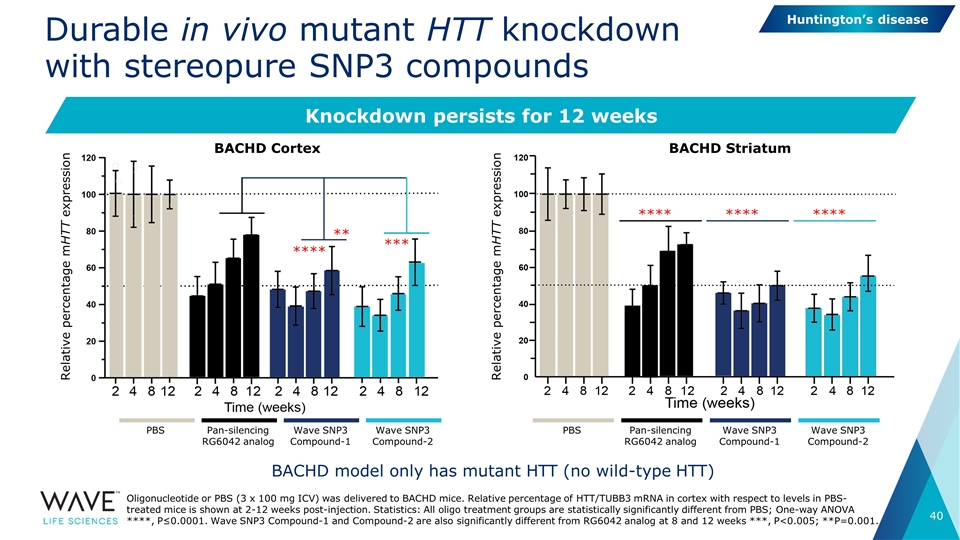
Durable in vivo mutant HTT knockdown with stereopure SNP3 compounds Knockdown persists for 12 weeks Oligonucleotide or PBS (3 x 100 mg ICV) was delivered to BACHD mice. Relative percentage of HTT/TUBB3 mRNA in cortex with respect to levels in PBS-treated mice is shown at 2-12 weeks post-injection. Statistics: All oligo treatment groups are statistically significantly different from PBS; One-way ANOVA ****, P≤0.0001. Wave SNP3 Compound-1 and Compound-2 are also significantly different from RG6042 analog at 8 and 12 weeks ***, P<0.005; **P=0.001. Relative percentage mHTT expression Relative percentage mHTT expression BACHD model only has mutant HTT (no wild-type HTT) PBS Pan-silencing RG6042 analog Wave SNP3 Compound-1 Wave SNP3 Compound-2 PBS Pan-silencing RG6042 analog Wave SNP3 Compound-1 Wave SNP3 Compound-2 BACHD Cortex BACHD Striatum ** **** *** **** **** **** Huntington’s disease

C9orf72 program Amyotrophic Lateral Sclerosis (ALS) Frontotemporal Dementia (FTD)
C9orf72: a critical genetic risk factor C9orf72 gene provides instructions for making protein found in various tissues, with abundance in nerve cells in the cerebral cortex and motor neurons C9orf72 genetic mutations are the strongest genetic risk factor found to date for the more common, non-inherited (sporadic) forms of Amyotrophic Lateral Sclerosis (ALS) and Frontotemporal Dementia (FTD); GGGGCC repeat drives the formation and accumulation of dipeptide repeat proteins that accumulate in brain tissue First pathogenic mechanism identified to be a genetic link between familial (inherited) ALS and FTD Most common mutation identified associated with familial ALS and FTD Availability of dipeptide biomarker in CSF has potential to accelerate drug development expanded GGGGCC repeat hexanucleotide repeat transcript Neuro C9orf72 Source: DeJesus-Hernandez M, Mackenzie IR, Boeve BF, et al. Neuron. 2011;72:245-256. Renton AE, Majounie E, Waite A, et al. Neuron. 2011;72:257-268.
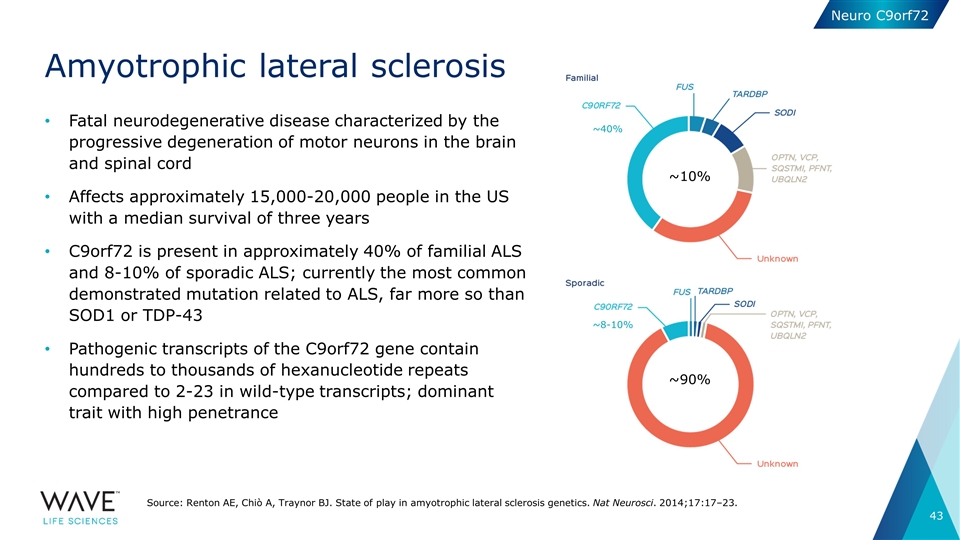
Amyotrophic lateral sclerosis Fatal neurodegenerative disease characterized by the progressive degeneration of motor neurons in the brain and spinal cord Affects approximately 15,000-20,000 people in the US with a median survival of three years C9orf72 is present in approximately 40% of familial ALS and 8-10% of sporadic ALS; currently the most common demonstrated mutation related to ALS, far more so than SOD1 or TDP-43 Pathogenic transcripts of the C9orf72 gene contain hundreds to thousands of hexanucleotide repeats compared to 2-23 in wild-type transcripts; dominant trait with high penetrance Source: Renton AE, Chiò A, Traynor BJ. State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci. 2014;17:17–23. Neuro C9orf72 ~40% ~8-10% ~10% ~90%

Frontotemporal dementia Progressive neuronal atrophy with loss in the frontal and temporal cortices characterized by personality and behavioral changes, as well as gradual impairment of language skills Affects approximately 55,000 people in the US Second most common form of early-onset dementia after Alzheimer’s disease in people under the age of 65 Up to 50% of FTD patients have a family history of dementia, many inheriting FTD as an autosomal dominant trait with high penetrance Pathogenic transcripts of the C9orf72 gene contain hundreds to thousands of hexanucleotide repeats compared to 2-23 in wild-type transcripts Neuro C9orf72 ~38% ~6% Sources: Stevens M, et al. Familial aggregation in frontotemporal dementia. Neurology. 1998;50:1541-1545. Majounie E, et al. Frequency of the C9orf72 hexanucleotide repeat expansion in patients with amyotrophic lateral sclerosis and frontotemporal dementia: a cross-sectional study. Lancet Neurol. 2012;11:323-330. 10% - 50% 50% - 90%
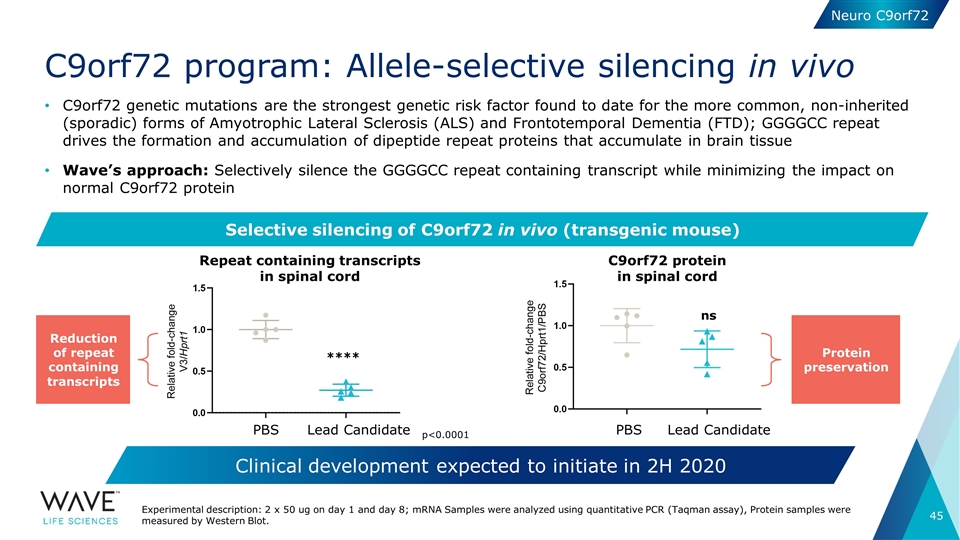
C9orf72 program: Allele-selective silencing in vivo C9orf72 genetic mutations are the strongest genetic risk factor found to date for the more common, non-inherited (sporadic) forms of Amyotrophic Lateral Sclerosis (ALS) and Frontotemporal Dementia (FTD); GGGGCC repeat drives the formation and accumulation of dipeptide repeat proteins that accumulate in brain tissue Wave’s approach: Selectively silence the GGGGCC repeat containing transcript while minimizing the impact on normal C9orf72 protein Repeat containing transcripts in spinal cord **** ns PBS Lead Candidate PBS Lead Candidate Experimental description: 2 x 50 ug on day 1 and day 8; mRNA Samples were analyzed using quantitative PCR (Taqman assay), Protein samples were measured by Western Blot. Selective silencing of C9orf72 in vivo (transgenic mouse) Clinical development expected to initiate in 2H 2020 C9orf72 protein in spinal cord Reduction of repeat containing transcripts Protein preservation p<0.0001 Neuro C9orf72

Ophthalmology
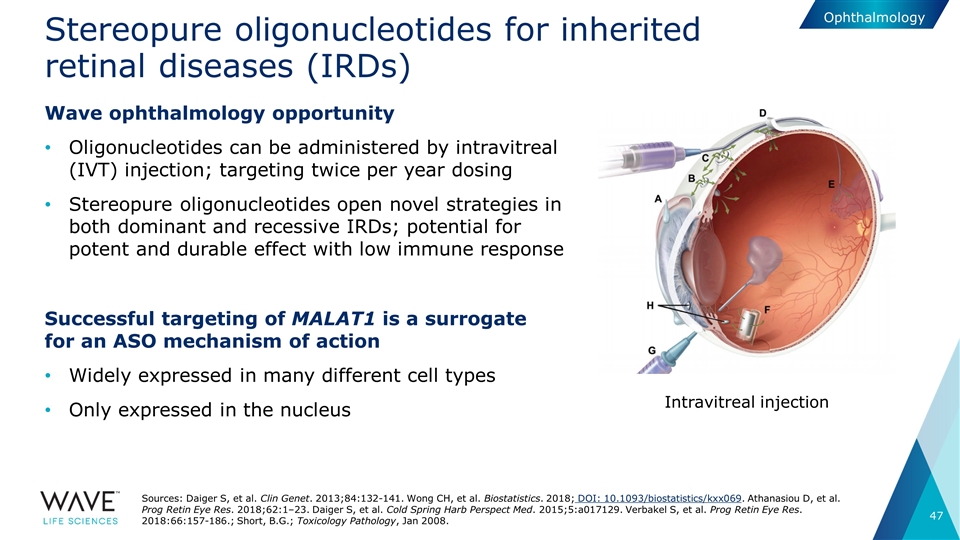
Stereopure oligonucleotides for inherited retinal diseases (IRDs) Wave ophthalmology opportunity Oligonucleotides can be administered by intravitreal (IVT) injection; targeting twice per year dosing Stereopure oligonucleotides open novel strategies in both dominant and recessive IRDs; potential for potent and durable effect with low immune response Successful targeting of MALAT1 is a surrogate for an ASO mechanism of action Widely expressed in many different cell types Only expressed in the nucleus Intravitreal injection Sources: Daiger S, et al. Clin Genet. 2013;84:132-141. Wong CH, et al. Biostatistics. 2018; DOI: 10.1093/biostatistics/kxx069. Athanasiou D, et al. Prog Retin Eye Res. 2018;62:1–23. Daiger S, et al. Cold Spring Harb Perspect Med. 2015;5:a017129. Verbakel S, et al. Prog Retin Eye Res. 2018:66:157-186.; Short, B.G.; Toxicology Pathology, Jan 2008. Ophthalmology
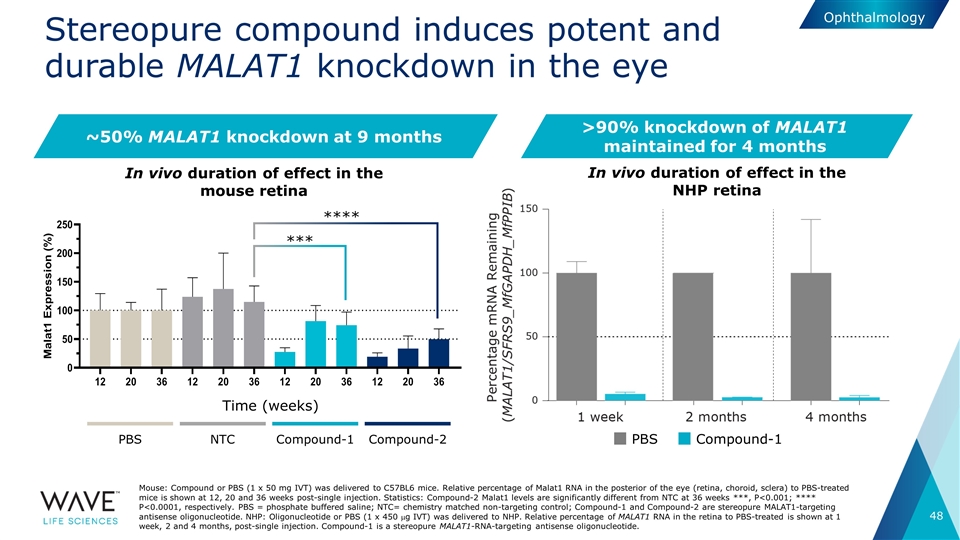
Stereopure compound induces potent and durable MALAT1 knockdown in the eye ~50% MALAT1 knockdown at 9 months Mouse: Compound or PBS (1 x 50 mg IVT) was delivered to C57BL6 mice. Relative percentage of Malat1 RNA in the posterior of the eye (retina, choroid, sclera) to PBS-treated mice is shown at 12, 20 and 36 weeks post-single injection. Statistics: Compound-2 Malat1 levels are significantly different from NTC at 36 weeks ***, P<0.001; **** P<0.0001, respectively. PBS = phosphate buffered saline; NTC= chemistry matched non-targeting control; Compound-1 and Compound-2 are stereopure MALAT1-targeting antisense oligonucleotide. NHP: Oligonucleotide or PBS (1 x 450 mg IVT) was delivered to NHP. Relative percentage of MALAT1 RNA in the retina to PBS-treated is shown at 1 week, 2 and 4 months, post-single injection. Compound-1 is a stereopure MALAT1-RNA-targeting antisense oligonucleotide. PBS NTC Compound-1 Compound-2 Time (weeks) **** *** Ophthalmology In vivo duration of effect in the NHP retina In vivo duration of effect in the mouse retina PBS Compound-1 >90% knockdown of MALAT1 maintained for 4 months
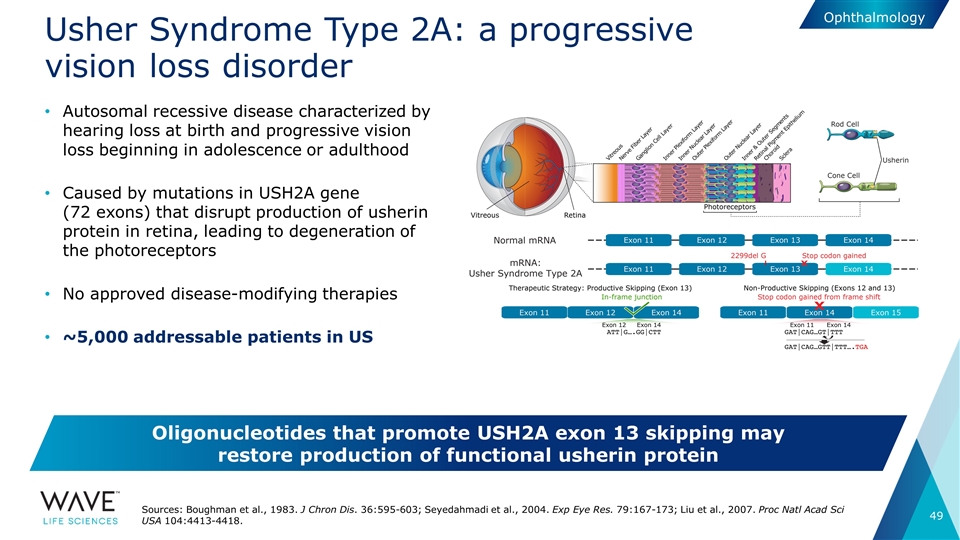
Usher Syndrome Type 2A: a progressive vision loss disorder Autosomal recessive disease characterized by hearing loss at birth and progressive vision loss beginning in adolescence or adulthood Caused by mutations in USH2A gene (72 exons) that disrupt production of usherin protein in retina, leading to degeneration of the photoreceptors No approved disease-modifying therapies ~5,000 addressable patients in US Sources: Boughman et al., 1983. J Chron Dis. 36:595-603; Seyedahmadi et al., 2004. Exp Eye Res. 79:167-173; Liu et al., 2007. Proc Natl Acad Sci USA 104:4413-4418. Oligonucleotides that promote USH2A exon 13 skipping may restore production of functional usherin protein Ophthalmology
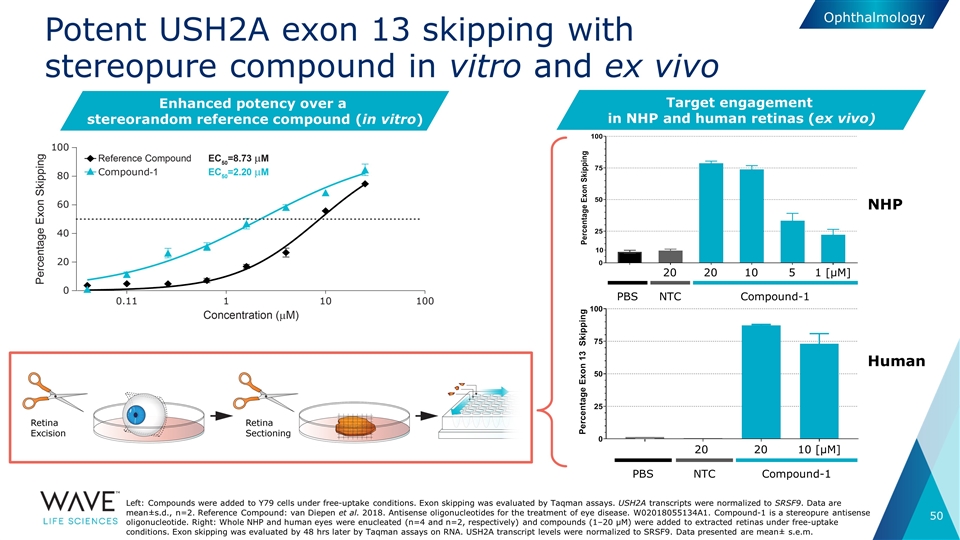
Potent USH2A exon 13 skipping with stereopure compound in vitro and ex vivo Left: Compounds were added to Y79 cells under free-uptake conditions. Exon skipping was evaluated by Taqman assays. USH2A transcripts were normalized to SRSF9. Data are mean±s.d., n=2. Reference Compound: van Diepen et al. 2018. Antisense oligonucleotides for the treatment of eye disease. W02018055134A1. Compound-1 is a stereopure antisense oligonucleotide. Right: Whole NHP and human eyes were enucleated (n=4 and n=2, respectively) and compounds (1–20 µM) were added to extracted retinas under free-uptake conditions. Exon skipping was evaluated by 48 hrs later by Taqman assays on RNA. USH2A transcript levels were normalized to SRSF9. Data presented are mean± s.e.m. Enhanced potency over a stereorandom reference compound (in vitro) Ophthalmology Target engagement in NHP and human retinas (ex vivo) PBS NTC Compound-1 20 20 10 5 1 [µM] PBS NTC Compound-1 20 20 10 [µM] NHP Human
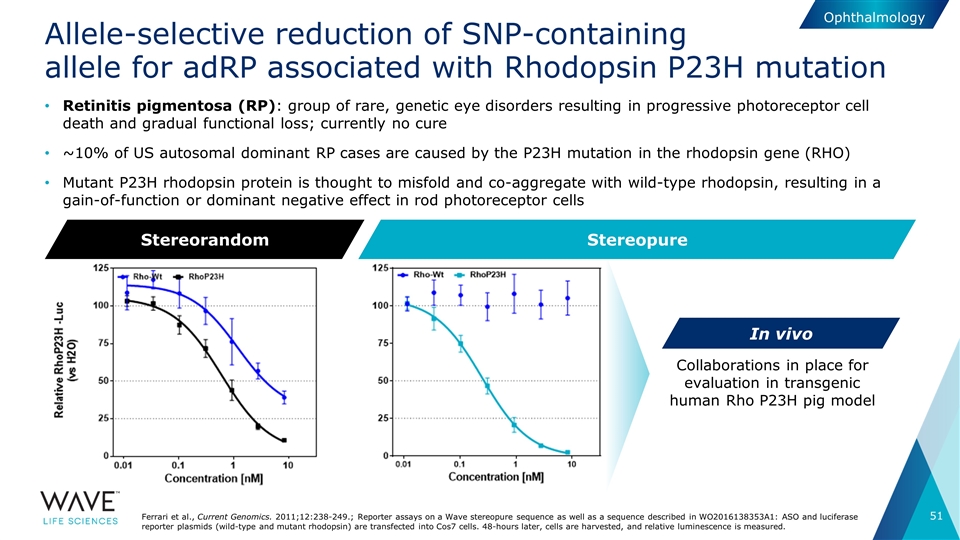
Allele-selective reduction of SNP-containing allele for adRP associated with Rhodopsin P23H mutation Ferrari et al., Current Genomics. 2011;12:238-249.; Reporter assays on a Wave stereopure sequence as well as a sequence described in WO2016138353A1: ASO and luciferase reporter plasmids (wild-type and mutant rhodopsin) are transfected into Cos7 cells. 48-hours later, cells are harvested, and relative luminescence is measured. Stereorandom Stereopure Collaborations in place for evaluation in transgenic human Rho P23H pig model In vivo Ophthalmology Retinitis pigmentosa (RP): group of rare, genetic eye disorders resulting in progressive photoreceptor cell death and gradual functional loss; currently no cure ~10% of US autosomal dominant RP cases are caused by the P23H mutation in the rhodopsin gene (RHO) Mutant P23H rhodopsin protein is thought to misfold and co-aggregate with wild-type rhodopsin, resulting in a gain-of-function or dominant negative effect in rod photoreceptor cells

ADAR-mediated RNA editing
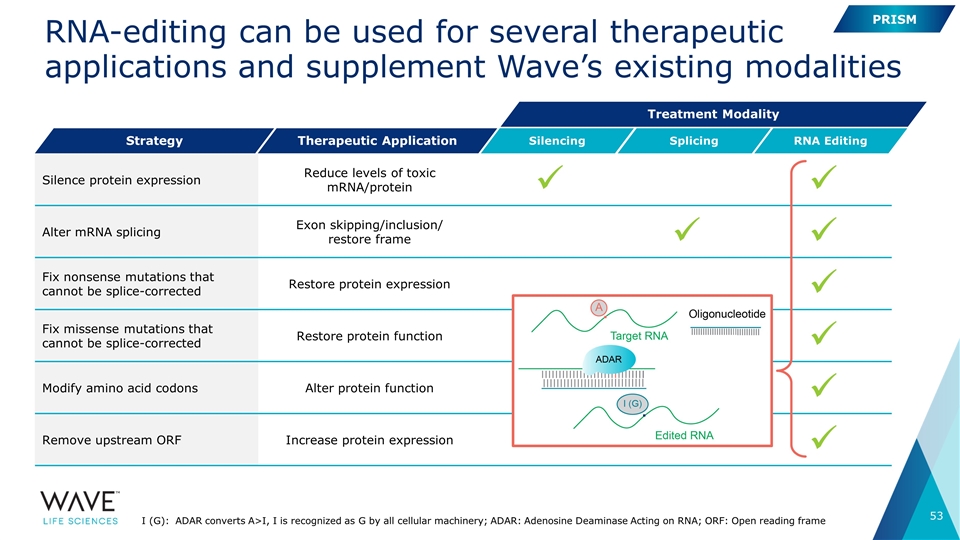
RNA-editing can be used for several therapeutic applications and supplement Wave’s existing modalities Silence protein expression Reduce levels of toxic mRNA/protein ü ü Alter mRNA splicing Exon skipping/inclusion/ restore frame ü ü Fix nonsense mutations that cannot be splice-corrected Restore protein expression ü Fix missense mutations that cannot be splice-corrected Restore protein function ü Modify amino acid codons Alter protein function ü Remove upstream ORF Increase protein expression ü Strategy Therapeutic Application Silencing Splicing RNA Editing Treatment Modality PRISM I (G): ADAR converts A>I, I is recognized as G by all cellular machinery; ADAR: Adenosine Deaminase Acting on RNA; ORF: Open reading frame
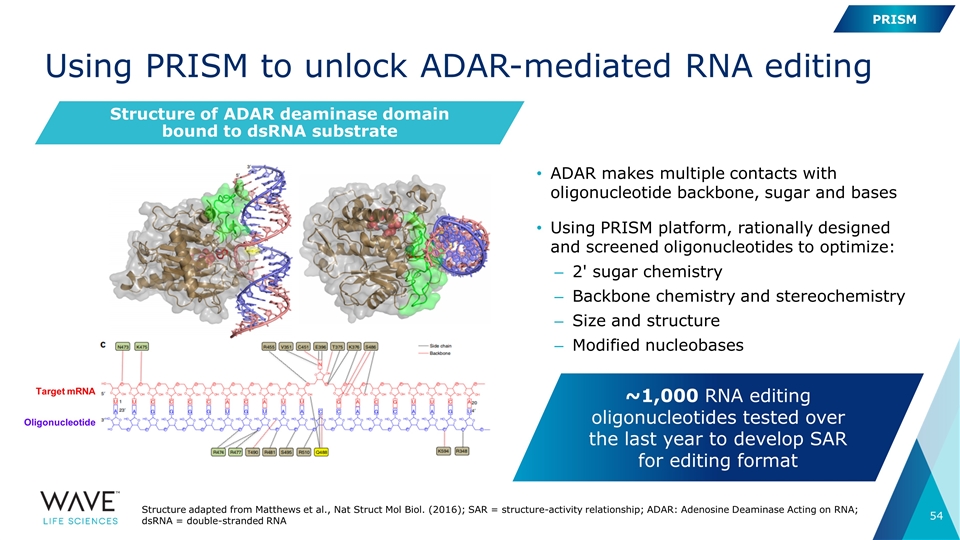
Structure of ADAR deaminase domain bound to dsRNA substrate Using PRISM to unlock ADAR-mediated RNA editing Target mRNA Oligonucleotide ADAR makes multiple contacts with oligonucleotide backbone, sugar and bases Using PRISM platform, rationally designed and screened oligonucleotides to optimize: 2' sugar chemistry Backbone chemistry and stereochemistry Size and structure Modified nucleobases ~1,000 RNA editing oligonucleotides tested over the last year to develop SAR for editing format PRISM Structure adapted from Matthews et al., Nat Struct Mol Biol. (2016); SAR = structure-activity relationship; ADAR: Adenosine Deaminase Acting on RNA; dsRNA = double-stranded RNA
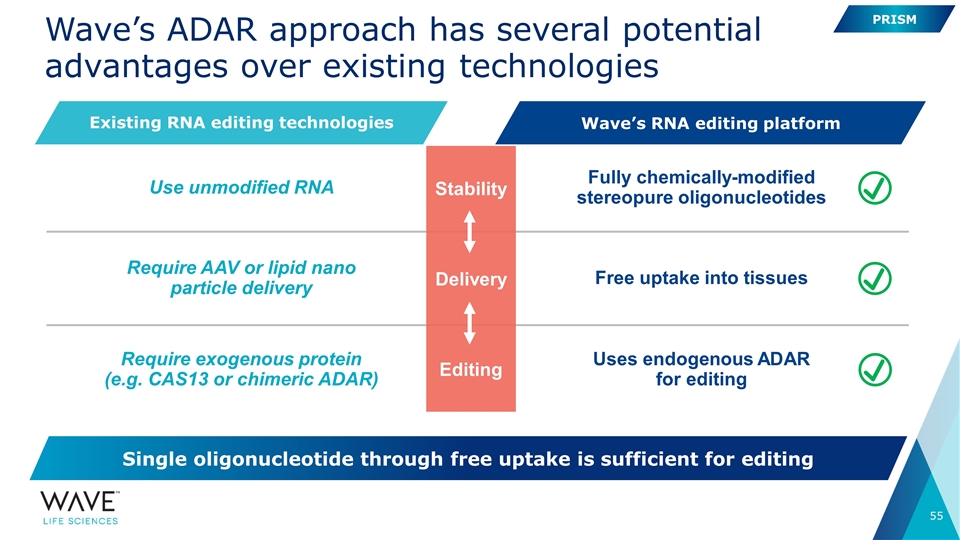
Wave��s ADAR approach has several potential advantages over existing technologies Require AAV or lipid nano particle delivery Require exogenous protein (e.g. CAS13 or chimeric ADAR) Free uptake into tissues Uses endogenous ADAR for editing Use unmodified RNA Fully chemically-modified stereopure oligonucleotides Stability Delivery Editing PRISM Existing RNA editing technologies Wave’s RNA editing platform Single oligonucleotide through free uptake is sufficient for editing
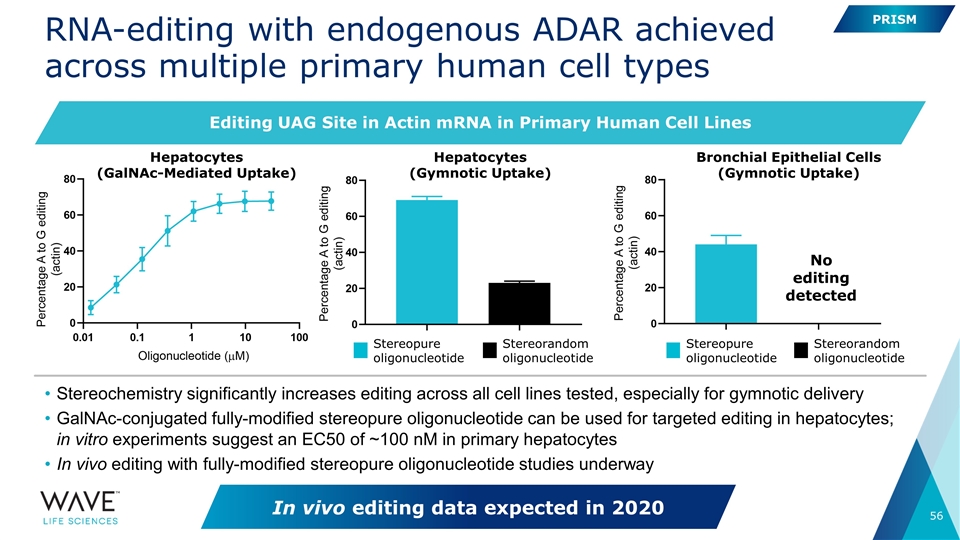

RNA-editing with endogenous ADAR achieved across multiple primary human cell types Stereochemistry significantly increases editing across all cell lines tested, especially for gymnotic delivery GalNAc-conjugated fully-modified stereopure oligonucleotide can be used for targeted editing in hepatocytes; in vitro experiments suggest an EC50 of ~100 nM in primary hepatocytes In vivo editing with fully-modified stereopure oligonucleotide studies underway Hepatocytes (GalNAc-Mediated Uptake) Hepatocytes (Gymnotic Uptake) Bronchial Epithelial Cells (Gymnotic Uptake) Stereopure oligonucleotide Stereorandom oligonucleotide No editing detected Stereopure oligonucleotide Stereorandom oligonucleotide PRISM In vivo editing data expected in 2020 Editing UAG Site in Actin mRNA in Primary Human Cell Lines

Anticipated upcoming Wave milestones Neuromuscular 4Q 2019: Interim dystrophin data readout for suvodirsen from OLE in DMD (exon 51) 2H 2020: Accelerated approval filing for suvodirsen in DMD (exon 51) in US, pending positive clinical dystrophin expression data 2H 2020: Topline clinical data for WVE-N531 in DMD (exon 53) CNS By YE 2019: Topline data readout from PRECISION-HD2 Phase 1b/2a trial in Huntington’s disease Early 2020: Topline data readout from PRECISION-HD1 Phase 1b/2a trial in Huntington’s disease 2H 2020: Initiation of clinical development of C9orf72 program in ALS and FTD Ophthalmology 2020: Advance USH2A exon-skipping program RNA-editing 2020: In vivo ADAR editing data

Realizing the potential of genetic medicines For more information: Kate Rausch, Investor Relations krausch@wavelifesci.com 617.949.4827

























































