
From Serendipity to Rational Design Taking Molecular Glue Degraders to New Heights | October 2023

Forward-Looking Statements These materials include express and implied “forward-looking statements,” including forward-looking statements within the meaning of the Private Securities Litigation Reform Act of 1995. Forward looking statements include all statements that are not historical facts, and in some cases, can be identified by terms such as “may,” “might,” “will,” “could,” “would,” “should,” “expect,” “intend,” “plan,” “objective,” “anticipate,” “believe,” “estimate,” “predict,” “potential,” “continue,” “ongoing,” or the negative of these terms, or other comparable terminology intended to identify statements about the future. Forward-looking statements contained in herein include, but are not limited to, statements about our product development activities, including our expectations around the potential of molecular glue degraders (“MGDs”), the potential of our herein detailed pipeline and library of MGDs, including our MGDs for NEK7, CDK2, SCD targets, and our early stage undisclosed MGDs, our expectations for our collaboration with Roche, including the discovery and development of MGDs therefrom, our ongoing pre-clinical and clinical development of our GSPT1 degrader referred to as MRT-2359, including our expectations regarding the potential relevance of certain interim clinical data, our expectations for the nature and timing of any additional clinical data releases for MRT-2359, including any full phase 1 clinical data release, and our expectations for the nature and timing of our ongoing and future clinical development of MRT-2359, including our plan to continue the Phase 1/2 study of MRT-2359 and its anticipated timing and progress, and our ability to enroll the requisite number of patients and dose each dosing cohort in the intended manner, the potential for MRT-2359 to benefit patients living with a variety of difficult-to-treat cancers, our expectations regarding the advancement, and timing thereof, of our pipeline and the various products therein, our ability to advance our development candidates, including MRT-6160 and our expectations for MRT-6160 to enter the clinic in 2024 and its potential applications across multiple autoimmune diseases, our expectations regarding potential therapeutic opportunities for our MGDs, and our clinical development expectations therefor, our expectations regarding patient populations and medical needs for any potential therapeutic opportunities for our MGDs, our expectations regarding our proprietary QuEEN™ platform and its potential to be highly productive and an industry-leading platform for designing MGDs to address hard-to-drug disease targets via degradation, combining experimentation with AI to push the boundaries of what is possible with MGDs, the nature and our expectations for our rationally-designed library of MGDs, including our ability to continue its expansion, and its ability to reprogram ligases surfaces, unlock novel degrons and target space, and the strength of our financial position, including our estimates of cash runway, among others. By their nature, these statements are subject to numerous risks and uncertainties, as well as those risks and uncertainties set forth in our Annual Report on Form 10-K for the fourth quarter and full year ended December 31, 2022, filed, with the US Securities and Exchange Commission on March 16, 2023, and any subsequent filings, that could cause actual results, performance or achievement to differ materially and adversely from those anticipated or implied in the statements. You should not rely upon forward looking statements as predictions of future events. Although our management believes that the expectations reflected in our statements are reasonable, we cannot guarantee that the future results, performance or events and circumstances described in the forward-looking statements will be achieved or occur. Recipients are cautioned not to place undue reliance on these forward-looking statements, which speak only as of the date such statements are made and should not be construed as statements of fact. We undertake no obligation to publicly update any forward-looking statements, whether as a result of new information, any future presentations or otherwise, except as required by applicable law. Certain information contained in these materials and any statements made orally during any presentation of these materials that relate to the materials or are based on studies, publications, surveys and other data obtained from third-party sources and our own internal estimates and research. While we believe these third-party studies, publications, surveys and other data to be reliable as of the date of these materials, we have not independently verified, and make no representations as to the adequacy, fairness, accuracy or completeness of, any information obtained from third-party sources. In addition, no independent source has evaluated the reasonableness or accuracy of our internal estimates or research and no reliance should be made on any information or statements made in these materials relating to or based on such internal estimates and research. These materials remain the proprietary intellectual property of Monte Rosa Therapeutics and should not be distributed or reproduced in whole or in part without the prior written consent of Monte Rosa Therapeutics.

Monte Rosa Therapeutics - Highlights Taking molecular glue degraders (MGDs) to new heights Rationally designed MGDs with potential to solve many of the limitations of other modalities by selectively degrading therapeutically relevant proteins considered undruggable or inadequately drugged Highly productive, industry-leading platform for designing MGDs to address hard-to-drug disease targets via degradation, combining experimentation with AI to push the boundaries of what is possible with MGDs Five promising, wholly-owned programs spanning oncology, autoimmune, inflammation and other TAs Strong financial position providing cash runway into Q3 2025 Phase 1/2 clinical study ongoing with MRT-2359, with potential to treat difficult-to-drug MYC-driven cancers; optimal pharmacodynamic modulation and early signs of clinical activity observed Partnership with Roche to develop MGDs for oncology and neurological conditions – expands platform reach into neurology MRT-6160, highly selective VAV1-directed MGD, expected to enter clinic in mid-2024, with wide potential applications across autoimmune diseases

Molecular Glue Degraders (MGDs) - Drugging The Undruggable Expanding target space, fostering a new generation of drugs Drugging the Druggable INHIBITOR Drugging the Undruggable MGDs Redrugging the Druggable PROTAC UNDRUGGABLE TARGETS E3 Ligase DRUGGABLE TARGETS E3 Ligase DRUGGABLE TARGETS Target Space Expanding the Degradable Proteome
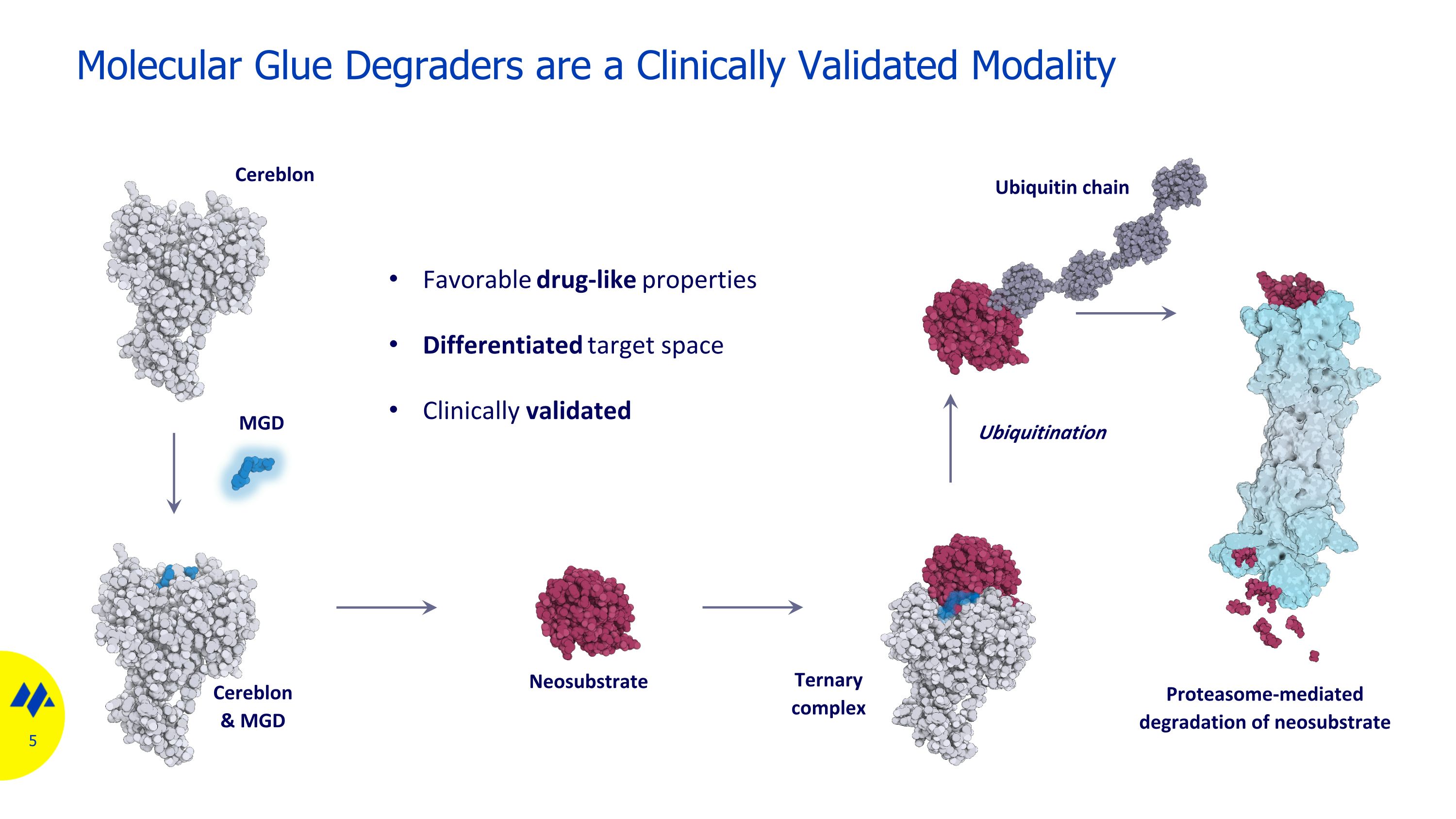

Molecular Glue Degraders are a Clinically Validated Modality MGD Cereblon Cereblon & MGD Neosubstrate Ternary complex Favorable drug-like properties Differentiated target space Clinically validated Ubiquitination Proteasome-mediated degradation of neosubstrate Ubiquitin chain
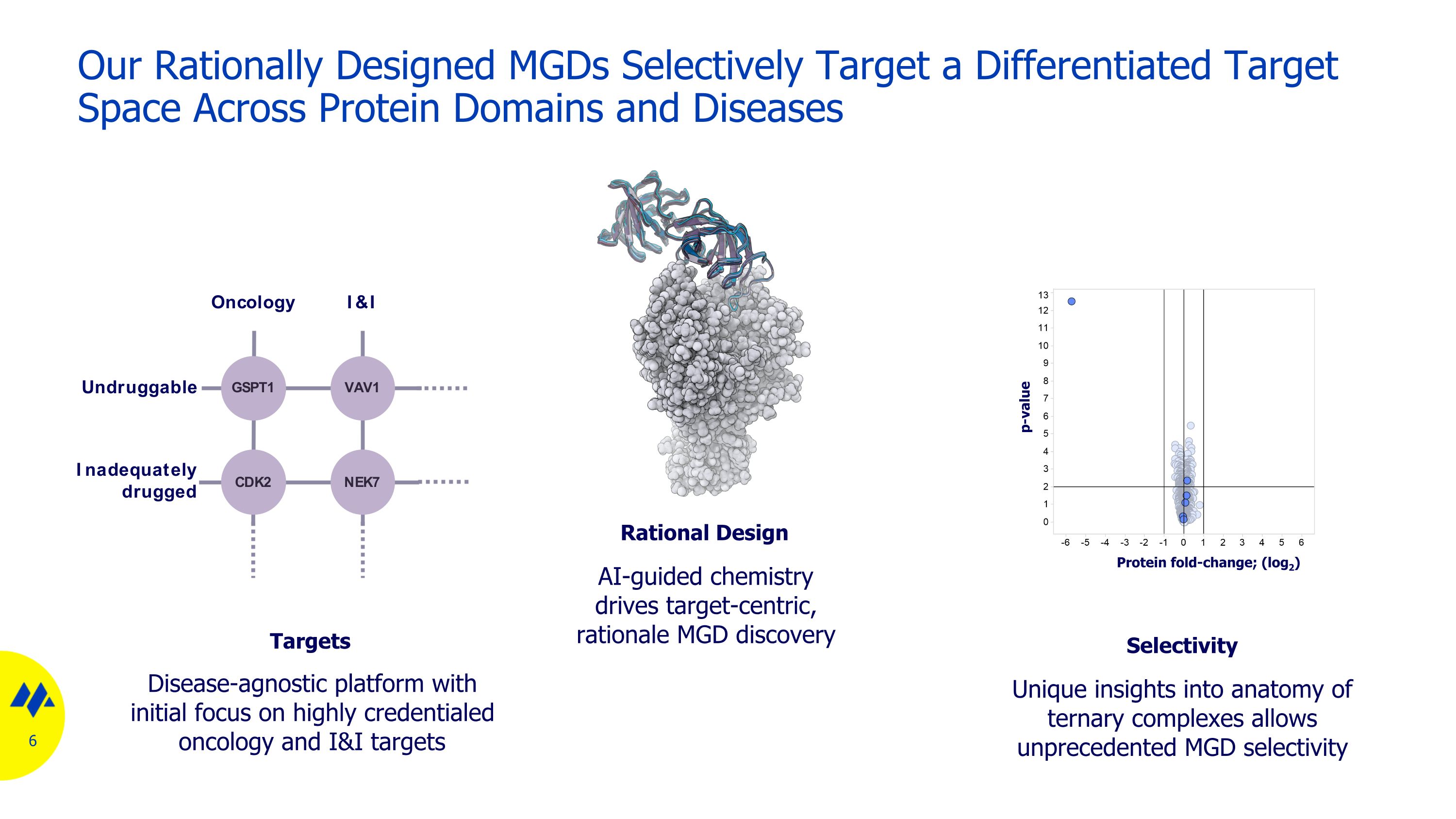
Our Rationally Designed MGDs Selectively Target a Differentiated Target Space Across Protein Domains and Diseases Unique insights into anatomy of ternary complexes allows unprecedented MGD selectivity Selectivity Protein fold-change; (log2) p-value Rational Design AI-guided chemistry drives target-centric, rationale MGD discovery Disease-agnostic platform with initial focus on highly credentialed oncology and I&I targets Targets

Monte Rosa Therapeutics – Key Firsts and Accomplishments From serendipity to rational design of MGDs Established a target-centric drug discovery approach combining experimentation with AI enabling rational design of highly potent and selective MGDs Progressed VAV1 MGD MRT-6160 into IND enabling studies; MRT-6160 is the first known MGD specifically developed for a non-oncology indication Built a proprietary molecular glue-based targeted protein degradation platform developing breakthrough therapeutics that selectively degrade disease-causing proteins Identified several additional highly credentialed targets as amenable to degradation through our platform including CDK2, NEK7 and multiple targets in SCD; began expanding approach to E3 ligases beyond Cereblon Presented interim data from Phase 1/2 trial of GSPT1-directed MGD MRT-2359 for the treatment of MYC-driven tumors; favorable safety profile and initial clinical signals observed – a first in solid tumors Established validating and broad discovery collaboration with Roche in oncology and neurological diseases

Portfolio
Targets Clinical Path Leveraging a Leading Drug Discovery Platform Purpose-built to discover and develop a wide landscape of therapeutically-relevant MGDs Undruggable and inadequately drugged proteins Proteins with non-catalytic and scaffolding functions High level of target validation, preclinically and clinically Programs with a biomarker-based patient selection strategy and a clear path to the clinic Opportunity for a rapid clinical PoC showing MOA and efficacy Address high unmet needs Drug a wide range of therapeutically-relevant proteins in oncology and beyond Create synergies within therapeutic areas Monte Rosa’s High-Value Proprietary Pipeline Patient Benefit
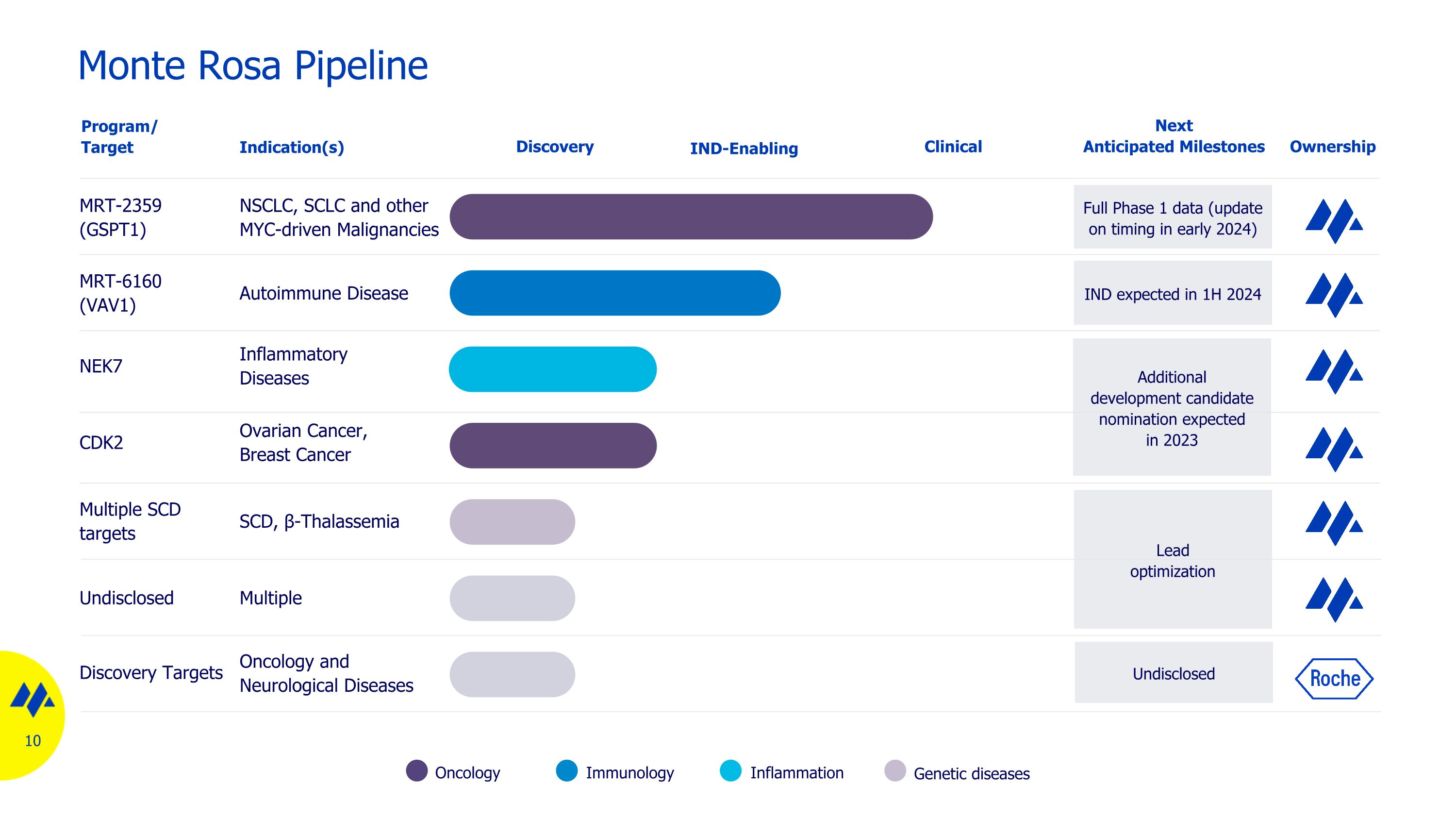
Monte Rosa Pipeline Oncology Inflammation Immunology Genetic diseases MRT-2359 (GSPT1) NSCLC, SCLC and other MYC-driven Malignancies NEK7 Inflammatory Diseases MRT-6160 (VAV1) Autoimmune Disease Discovery Program/ Target Indication(s) Full Phase 1 data (update on timing in early 2024) Multiple SCD targets SCD, β-Thalassemia Next Anticipated Milestones Ownership Undisclosed Multiple IND-Enabling Clinical Lead optimization Additional development candidate nomination expected in 2023 CDK2 Ovarian Cancer, Breast Cancer IND expected in 1H 2024 Discovery Targets Oncology and Neurological Diseases Undisclosed

GSPT1 program
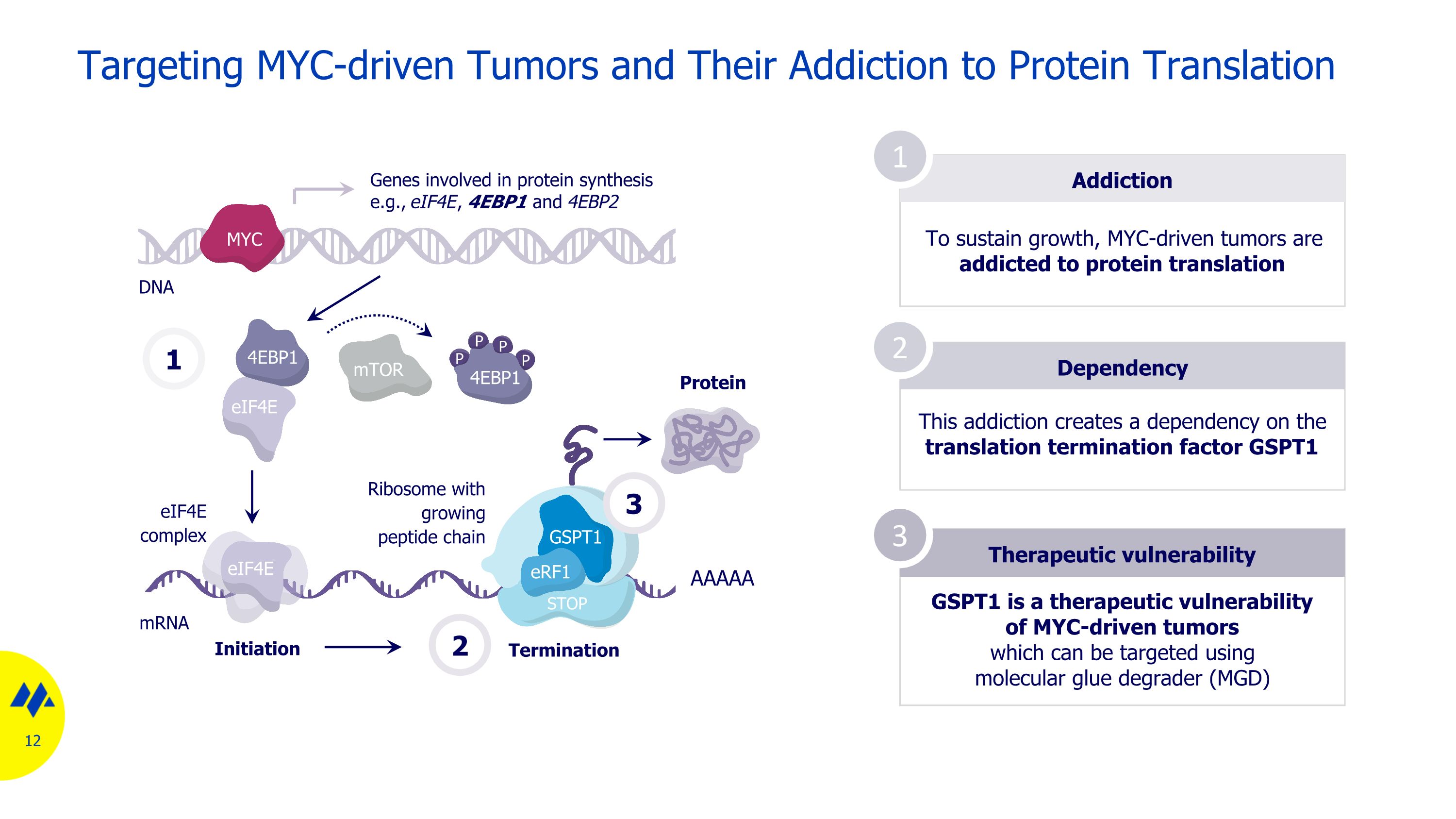
Targeting MYC-driven Tumors and Their Addiction to Protein Translation Addiction To sustain growth, MYC-driven tumors are addicted to protein translation Dependency Therapeutic vulnerability 1 2 3 This addiction creates a dependency on the translation termination factor GSPT1 GSPT1 is a therapeutic vulnerability of MYC-driven tumors which can be targeted using molecular glue degrader (MGD) mRNA DNA 1 mTOR eIF4E 4EBP1 P P P P 4EBP1 eIF4E eIF4E complex Genes involved in protein synthesis e.g., eIF4E, 4EBP1 and 4EBP2 Initiation Termination AAAAA Protein 2 MYC STOP GSPT1 eRF1 Ribosome with growing peptide chain 3
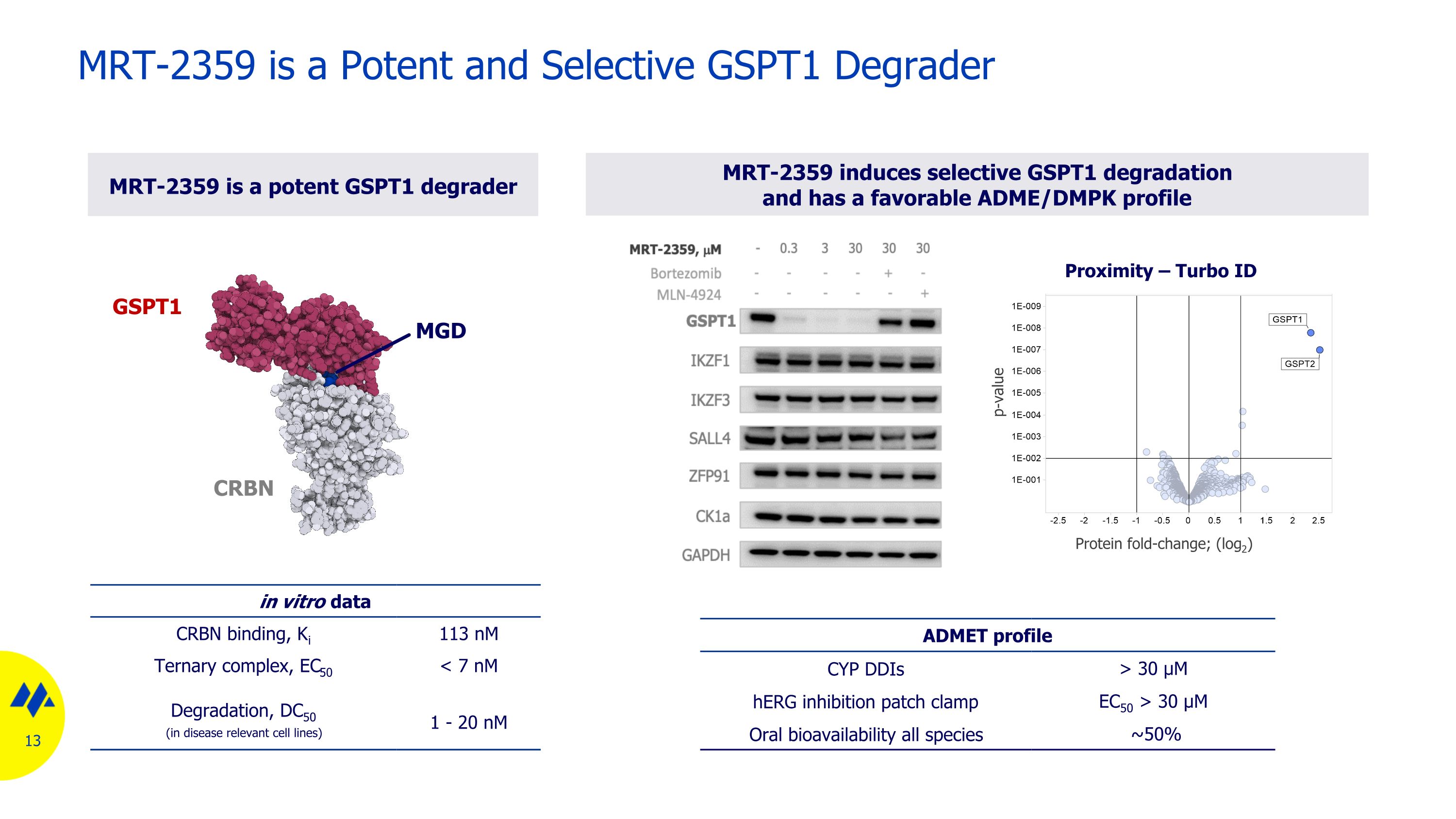
MRT-2359 is a Potent and Selective GSPT1 Degrader in vitro data CRBN binding, Ki 113 nM Ternary complex, EC50 < 7 nM Degradation, DC50 (in disease relevant cell lines) 1 - 20 nM MRT-2359 induces selective GSPT1 degradation and has a favorable ADME/DMPK profile MRT-2359 is a potent GSPT1 degrader ADMET profile CYP DDIs > 30 µM hERG inhibition patch clamp EC50 > 30 µM Oral bioavailability all species ~50% GSPT1 CRBN Protein fold-change; (log2) p-value Proximity – Turbo ID MGD
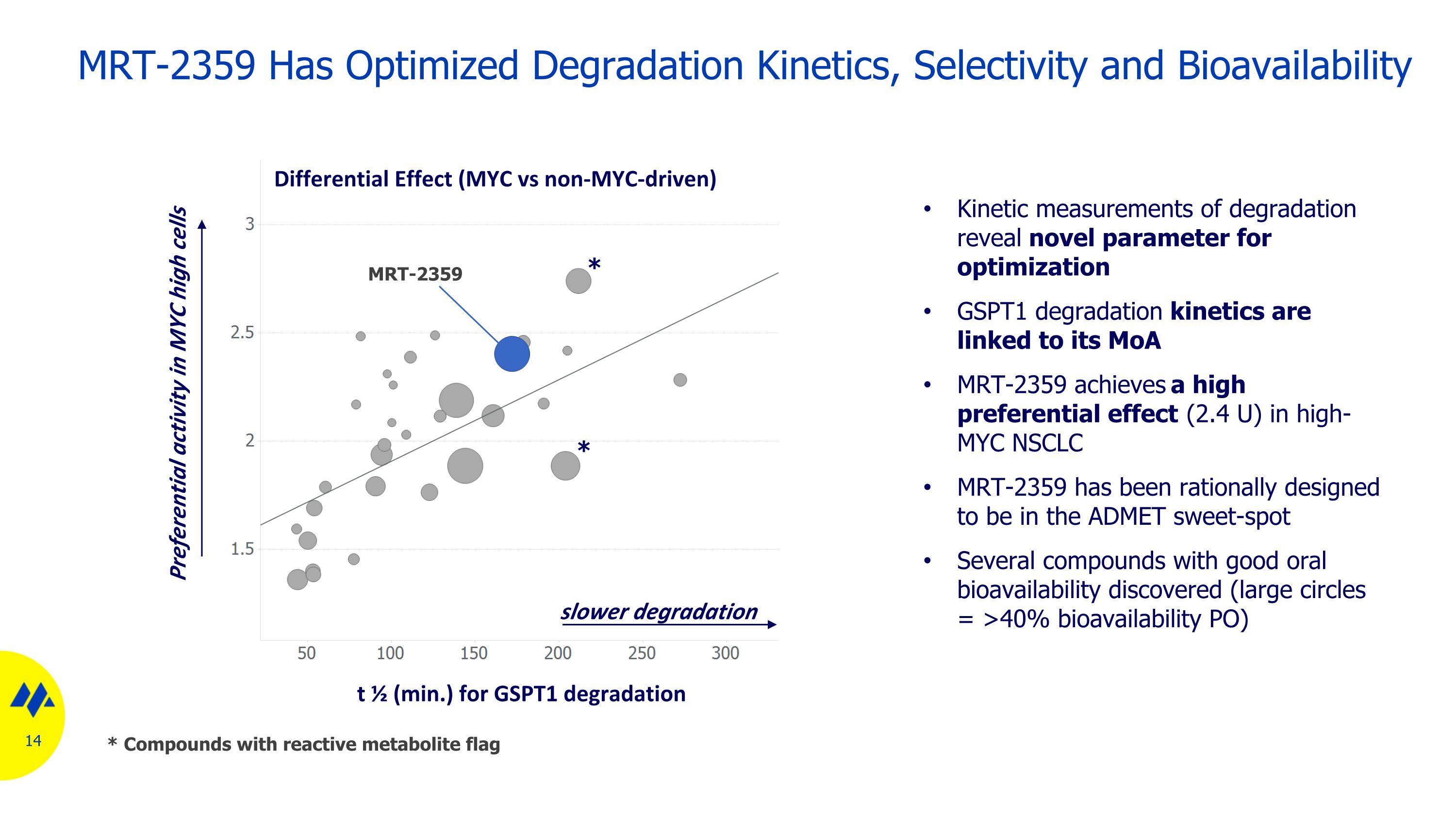
MRT-2359 Has Optimized Degradation Kinetics, Selectivity and Bioavailability Kinetic measurements of degradation reveal novel parameter for optimization GSPT1 degradation kinetics are linked to its MoA MRT-2359 achieves a high preferential effect (2.4 U) in high-MYC NSCLC MRT-2359 has been rationally designed to be in the ADMET sweet-spot Several compounds with good oral bioavailability discovered (large circles = >40% bioavailability PO) t ½ (min.) for GSPT1 degradation Differential Effect (MYC vs non-MYC-driven) slower degradation Preferential activity in MYC high cells MRT-2359 * * * Compounds with reactive metabolite flag
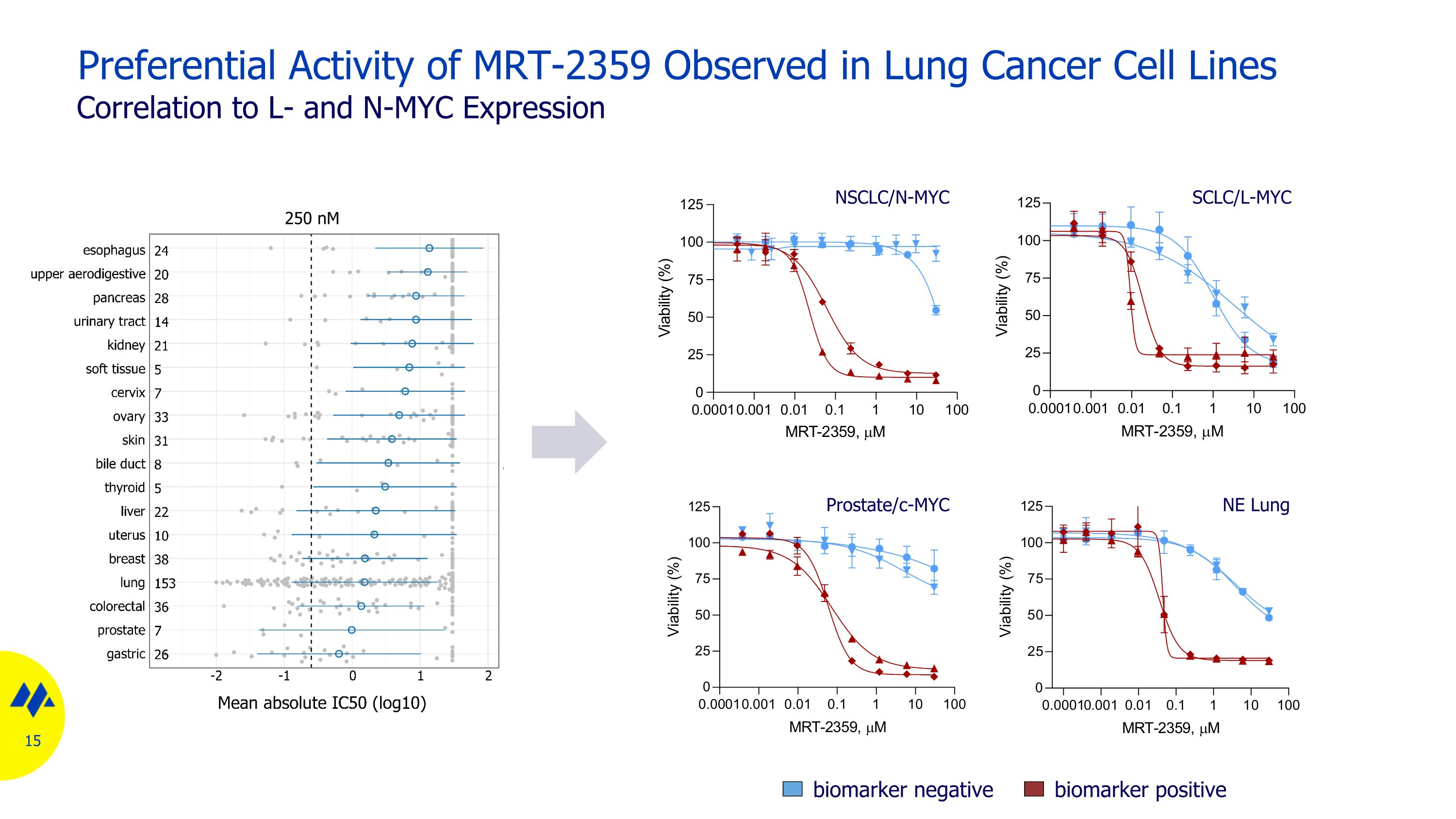
Preferential Activity of MRT-2359 Observed in Lung Cancer Cell Lines NSCLC/N-MYC Correlation to L- and N-MYC Expression SCLC/L-MYC NE Lung Prostate/c-MYC biomarker negative biomarker positive 250 nM
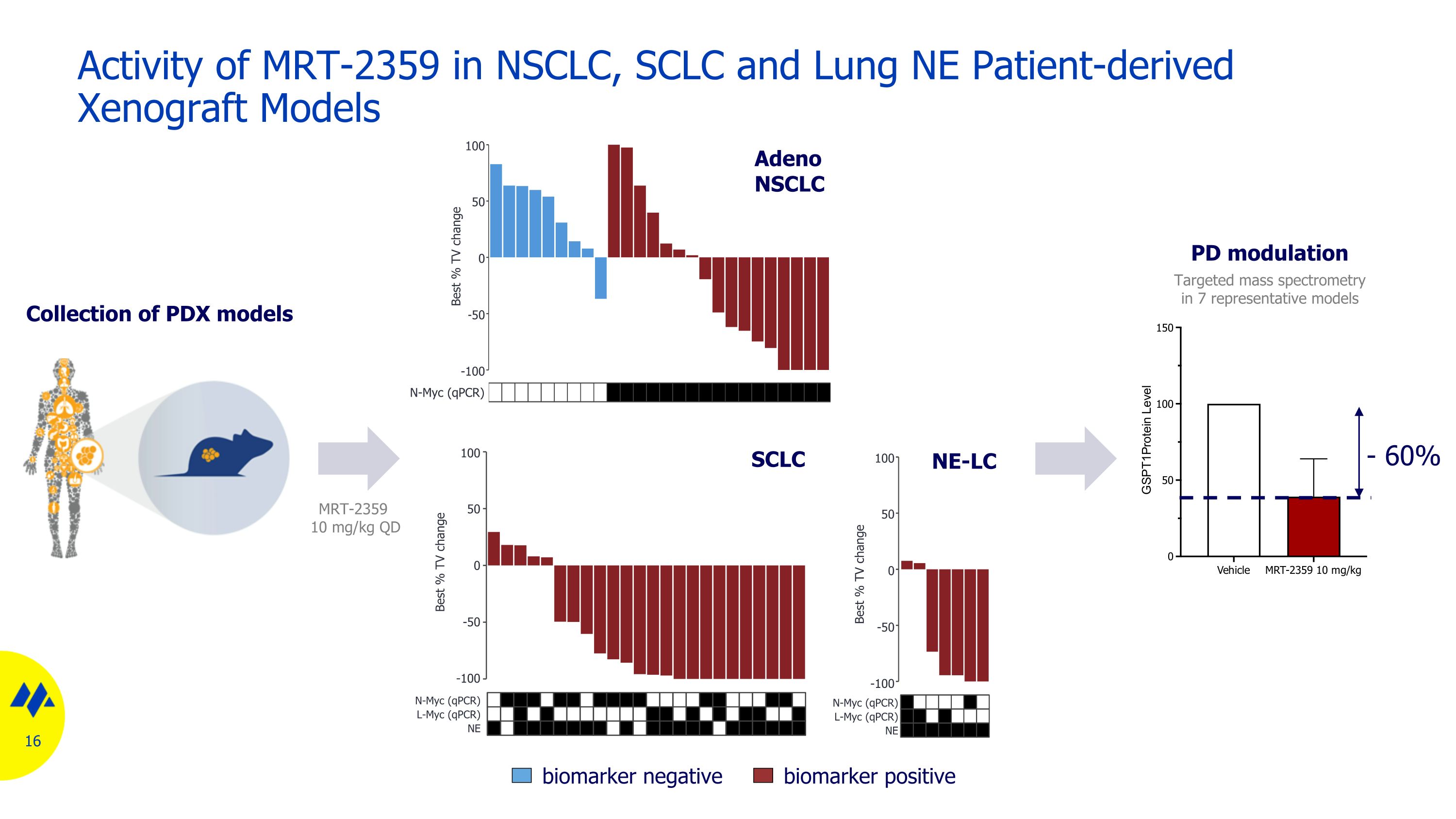
Activity of MRT-2359 in NSCLC, SCLC and Lung NE Patient-derived Xenograft Models Collection of PDX models 16 SCLC Adeno NSCLC NE-LC biomarker negative biomarker positive Targeted mass spectrometry in 7 representative models PD modulation 100 50 0 -50 -100 N-Myc (qPCR) Best % TV change 100 50 0 -50 -100 N-Myc (qPCR) Best % TV change L-Myc (qPCR) NE 100 50 0 -50 -100 N-Myc (qPCR) Best % TV change L-Myc (qPCR) NE MRT-2359 10 mg/kg QD - 60%
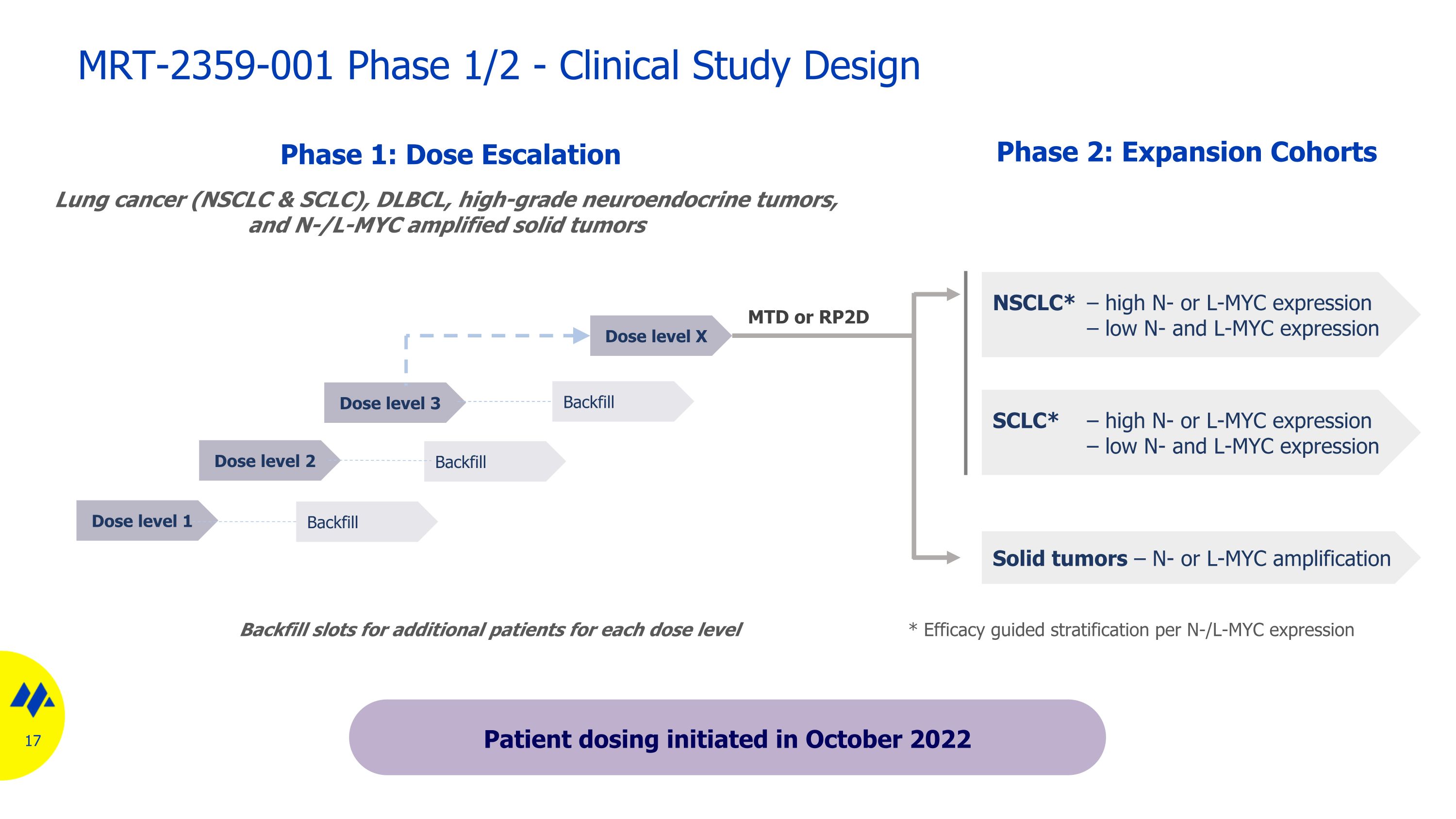
Dose level 1 MTD or RP2D Phase 2: Expansion Cohorts Phase 1: Dose Escalation NSCLC* – high N- or L-MYC expression – low N- and L-MYC expression Backfill slots for additional patients for each dose level Backfill Dose level 2 Dose level 3 Dose level X Backfill Backfill * Efficacy guided stratification per N-/L-MYC expression Solid tumors – N- or L-MYC amplification Lung cancer (NSCLC & SCLC), DLBCL, high-grade neuroendocrine tumors, and N-/L-MYC amplified solid tumors SCLC* – high N- or L-MYC expression – low N- and L-MYC expression MRT-2359-001 Phase 1/2 - Clinical Study Design Patient dosing initiated in October 2022

MRT-2359 Phase I Interim Data - October 2023
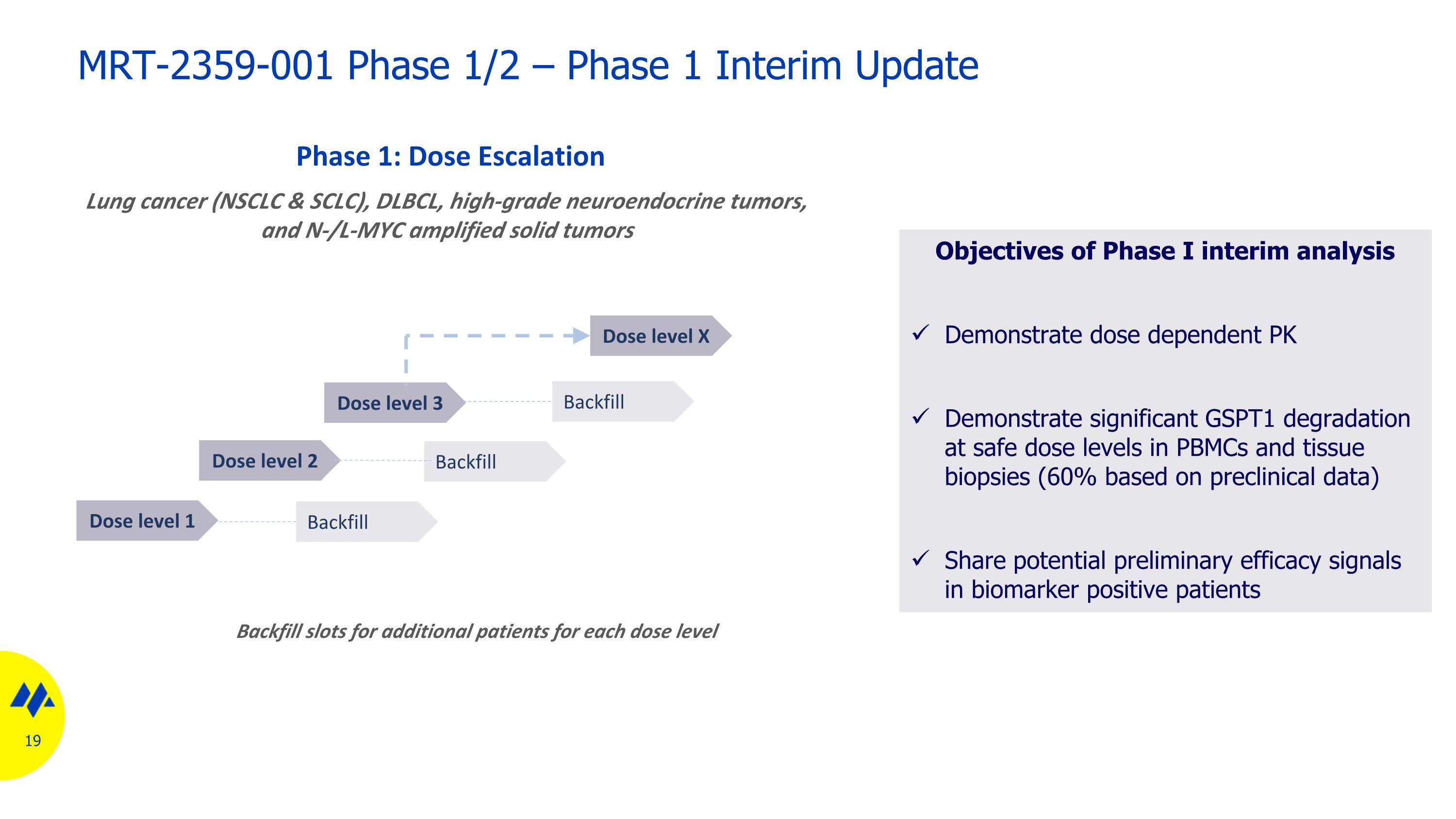
Dose level 1 Phase 1: Dose Escalation Backfill slots for additional patients for each dose level Backfill Dose level 2 Dose level 3 Dose level X Backfill Backfill Lung cancer (NSCLC & SCLC), DLBCL, high-grade neuroendocrine tumors, and N-/L-MYC amplified solid tumors MRT-2359-001 Phase 1/2 – Phase 1 Interim Update Objectives of Phase I interim analysis Demonstrate dose dependent PK Demonstrate significant GSPT1 degradation at safe dose levels in PBMCs and tissue biopsies (60% based on preclinical data) Share potential preliminary efficacy signals in biomarker positive patients

Executive Summary As of September 7th, 2023, 3 dose levels (0.5 mg, 1 mg, 2 mg) have been completed with 21 patients enrolled (including backfill patients) Observed dose dependent PK after oral dosing Clinical data support that 0.5 mg starting dose was fully active based on pharmacodynamic (PD) assessment of peripheral blood mononuclear cells (PBMC) and tissue samples from patients, with optimal PD modulation across dose levels tested Encouraging initial clinical activity at all dose levels: 2 partial responses (PRs) (1 confirmed, 1 unconfirmed) and 1 stable disease (SD) in 6 evaluable, heavily pretreated patients identified with biomarker positive tumors; 1 SD in non-small cell lung cancer (NSCLC) with un-evaluable biomarker status Safety profile supports continued development: Favorable adverse event (AE) profile at 0.5 and 1 mg Grade 4 thrombocytopenia identified as dose limiting toxicity (DLT) at 2 mg Previously reported limitations of CC-90009 not observed: No hypotension/cytokine release syndrome or clinically significant hypocalcemia observed at any dose level
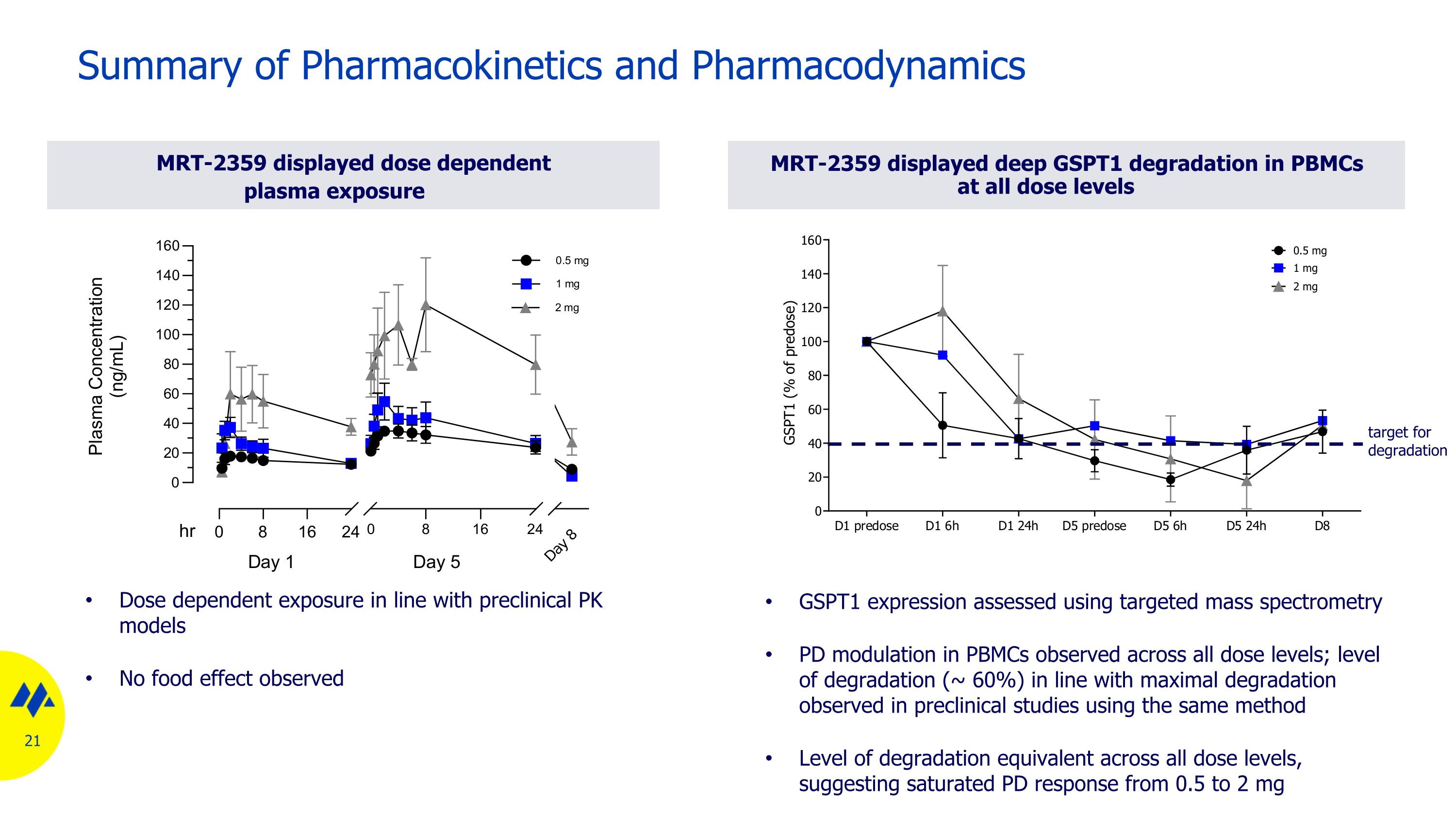
MRT-2359 displayed dose dependent plasma exposure Summary of Pharmacokinetics and Pharmacodynamics MRT-2359 displayed deep GSPT1 degradation in PBMCs at all dose levels GSPT1 expression assessed using targeted mass spectrometry PD modulation in PBMCs observed across all dose levels; level of degradation (~ 60%) in line with maximal degradation observed in preclinical studies using the same method Level of degradation equivalent across all dose levels, suggesting saturated PD response from 0.5 to 2 mg Dose dependent exposure in line with preclinical PK models No food effect observed target for degradation
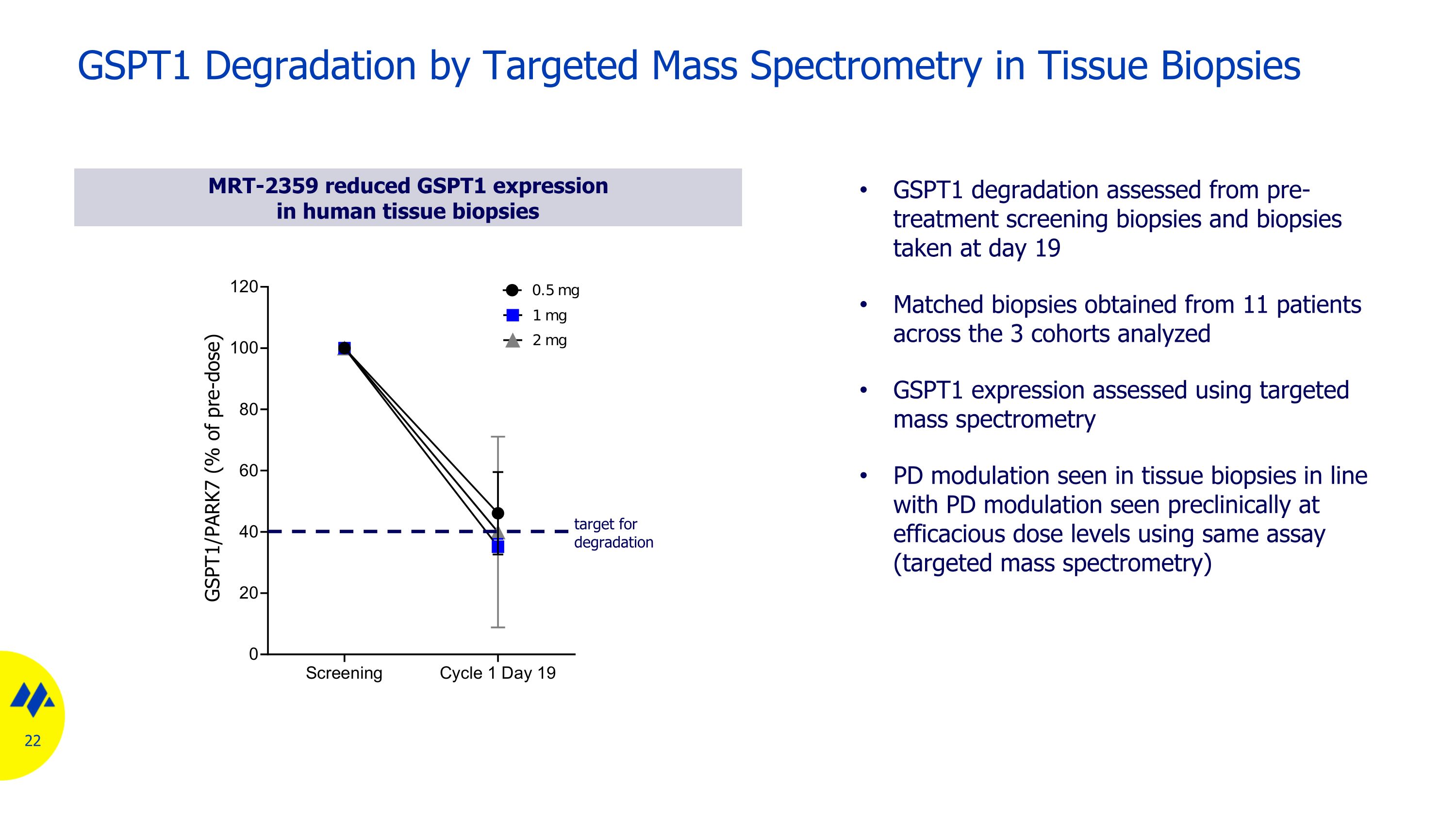
GSPT1 Degradation by Targeted Mass Spectrometry in Tissue Biopsies MRT-2359 reduced GSPT1 expression in human tissue biopsies GSPT1 degradation assessed from pre-treatment screening biopsies and biopsies taken at day 19 Matched biopsies obtained from 11 patients across the 3 cohorts analyzed GSPT1 expression assessed using targeted mass spectrometry PD modulation seen in tissue biopsies in line with PD modulation seen preclinically at efficacious dose levels using same assay (targeted mass spectrometry) target for degradation
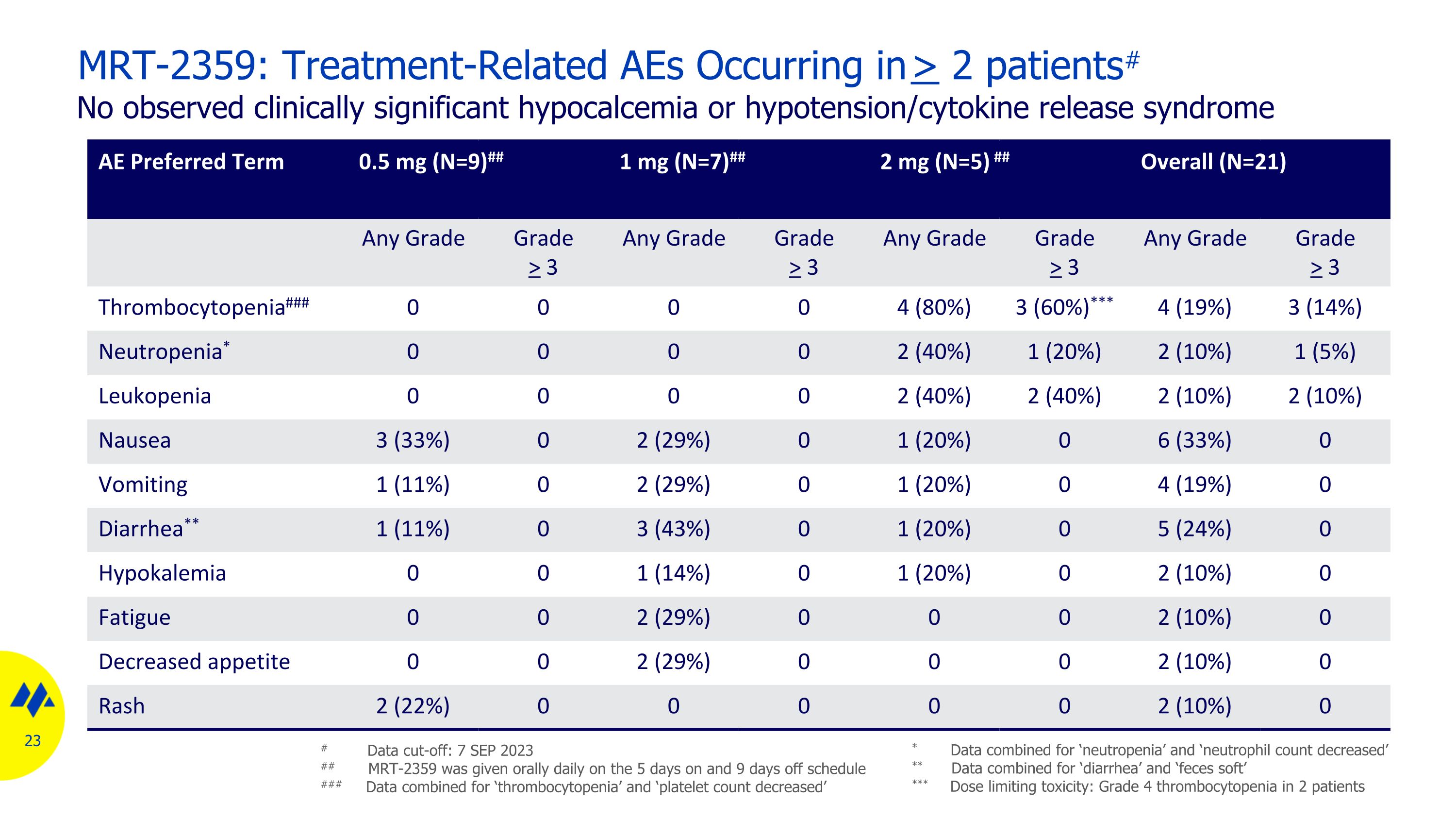
MRT-2359: Treatment-Related AEs Occurring in > 2 patients# No observed clinically significant hypocalcemia or hypotension/cytokine release syndrome AE Preferred Term 0.5 mg (N=9)## 1 mg (N=7)## 2 mg (N=5) ## Overall (N=21) Any Grade Grade > 3 Any Grade Grade > 3 Any Grade Grade > 3 Any Grade Grade > 3 Thrombocytopenia### 0 0 0 0 4 (80%) 3 (60%)*** 4 (19%) 3 (14%) Neutropenia* 0 0 0 0 2 (40%) 1 (20%) 2 (10%) 1 (5%) Leukopenia 0 0 0 0 2 (40%) 2 (40%) 2 (10%) 2 (10%) Nausea 3 (33%) 0 2 (29%) 0 1 (20%) 0 6 (33%) 0 Vomiting 1 (11%) 0 2 (29%) 0 1 (20%) 0 4 (19%) 0 Diarrhea** 1 (11%) 0 3 (43%) 0 1 (20%) 0 5 (24%) 0 Hypokalemia 0 0 1 (14%) 0 1 (20%) 0 2 (10%) 0 Fatigue 0 0 2 (29%) 0 0 0 2 (10%) 0 Decreased appetite 0 0 2 (29%) 0 0 0 2 (10%) 0 Rash 2 (22%) 0 0 0 0 0 2 (10%) 0 # Data cut-off: 7 SEP 2023 ## MRT-2359 was given orally daily on the 5 days on and 9 days off schedule ### Data combined for ‘thrombocytopenia’ and ‘platelet count decreased’ * Data combined for ‘neutropenia’ and ‘neutrophil count decreased’ ** Data combined for ‘diarrhea’ and ‘feces soft’ *** Dose limiting toxicity: Grade 4 thrombocytopenia in 2 patients
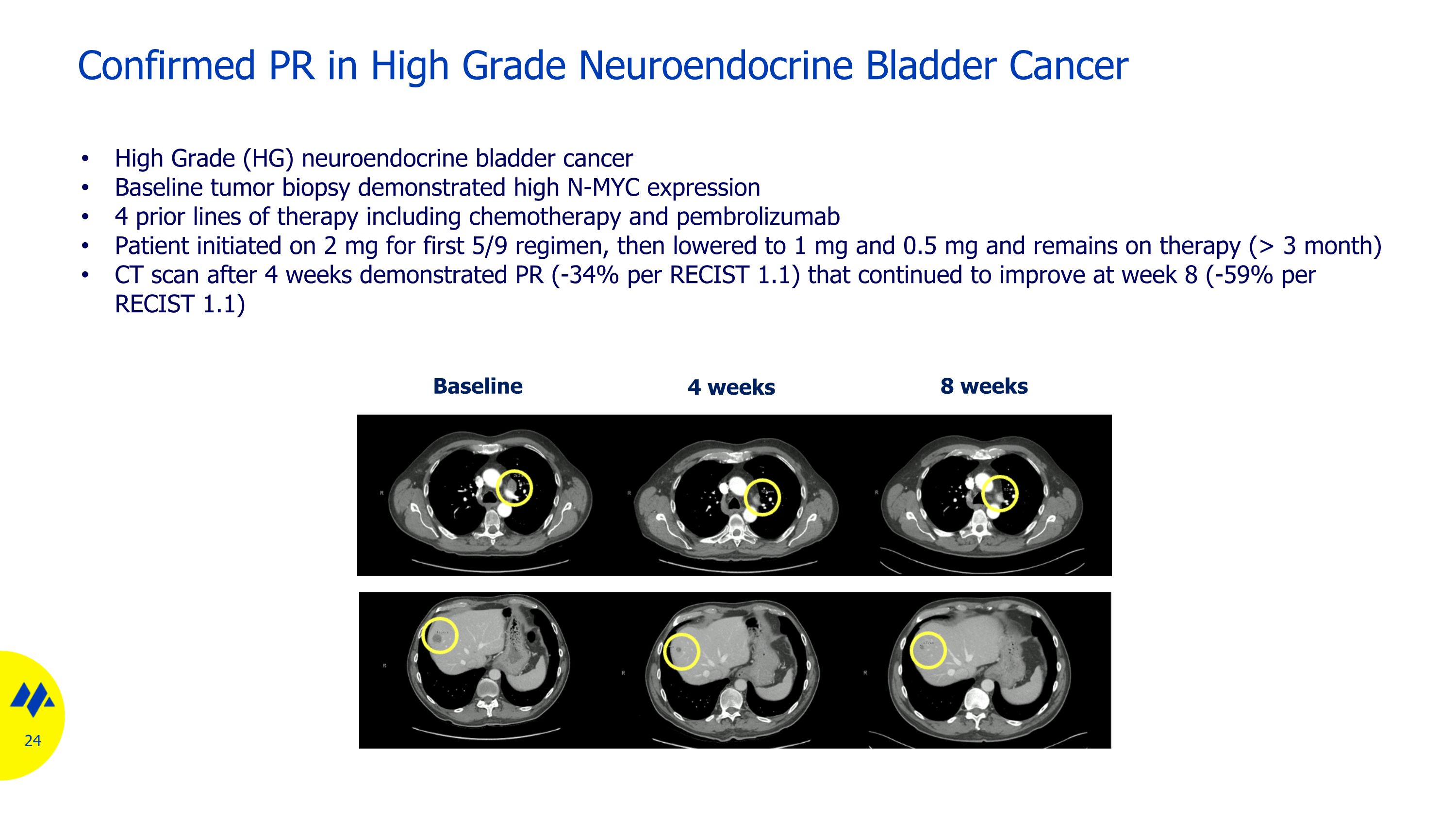
Confirmed PR in High Grade Neuroendocrine Bladder Cancer Baseline 8 weeks 4 weeks High Grade (HG) neuroendocrine bladder cancer Baseline tumor biopsy demonstrated high N-MYC expression 4 prior lines of therapy including chemotherapy and pembrolizumab Patient initiated on 2 mg for first 5/9 regimen, then lowered to 1 mg and 0.5 mg and remains on therapy (> 3 month) CT scan after 4 weeks demonstrated PR (-34% per RECIST 1.1) that continued to improve at week 8 (-59% per RECIST 1.1)
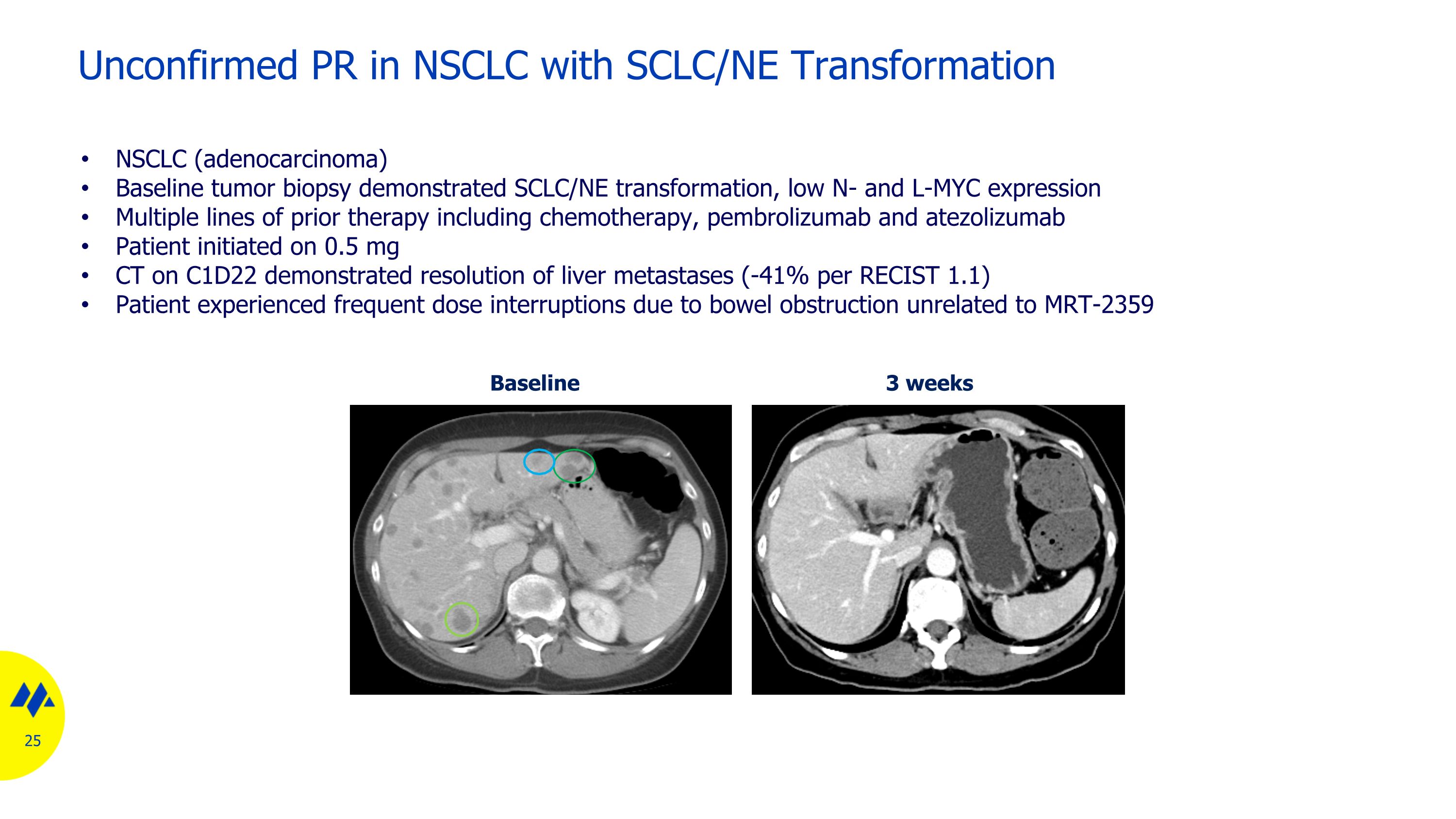
Unconfirmed PR in NSCLC with SCLC/NE Transformation Baseline 3 weeks NSCLC (adenocarcinoma) Baseline tumor biopsy demonstrated SCLC/NE transformation, low N- and L-MYC expression Multiple lines of prior therapy including chemotherapy, pembrolizumab and atezolizumab Patient initiated on 0.5 mg CT on C1D22 demonstrated resolution of liver metastases (-41% per RECIST 1.1) Patient experienced frequent dose interruptions due to bowel obstruction unrelated to MRT-2359

Summary and Next Steps Clinical data support that 0.5 mg starting dose was active based on PD assessment (PBMC and tissue), with optimal PD modulation across dose levels tested Encouraging initial clinical activity at all dose levels: 2 PRs (1 confirmed, 1 unconfirmed) and 1 SD in 6 evaluable, heavily pretreated patients identified with biomarker positive tumors; 1 SD in non-small cell lung cancer (NSCLC) with un-evaluable biomarker status Safety profile support further development: Favorable AE profile at 0.5 and 1 mg Grade 4 thrombocytopenia identified as dose limiting toxicity (DLT) at 2 mg Previously reported limitations of CC-90009 not observed: No hypotension/cytokine release syndrome or clinically significant hypocalcemia observed at any dose level Next Steps Currently dosing 1.5 mg, expected to complete DLT observation period by end of October Based on favorable tox profile, Company initiating intermittent dosing regimen of 21 days on and 7 days off drug (21/7)

CDK2 Program
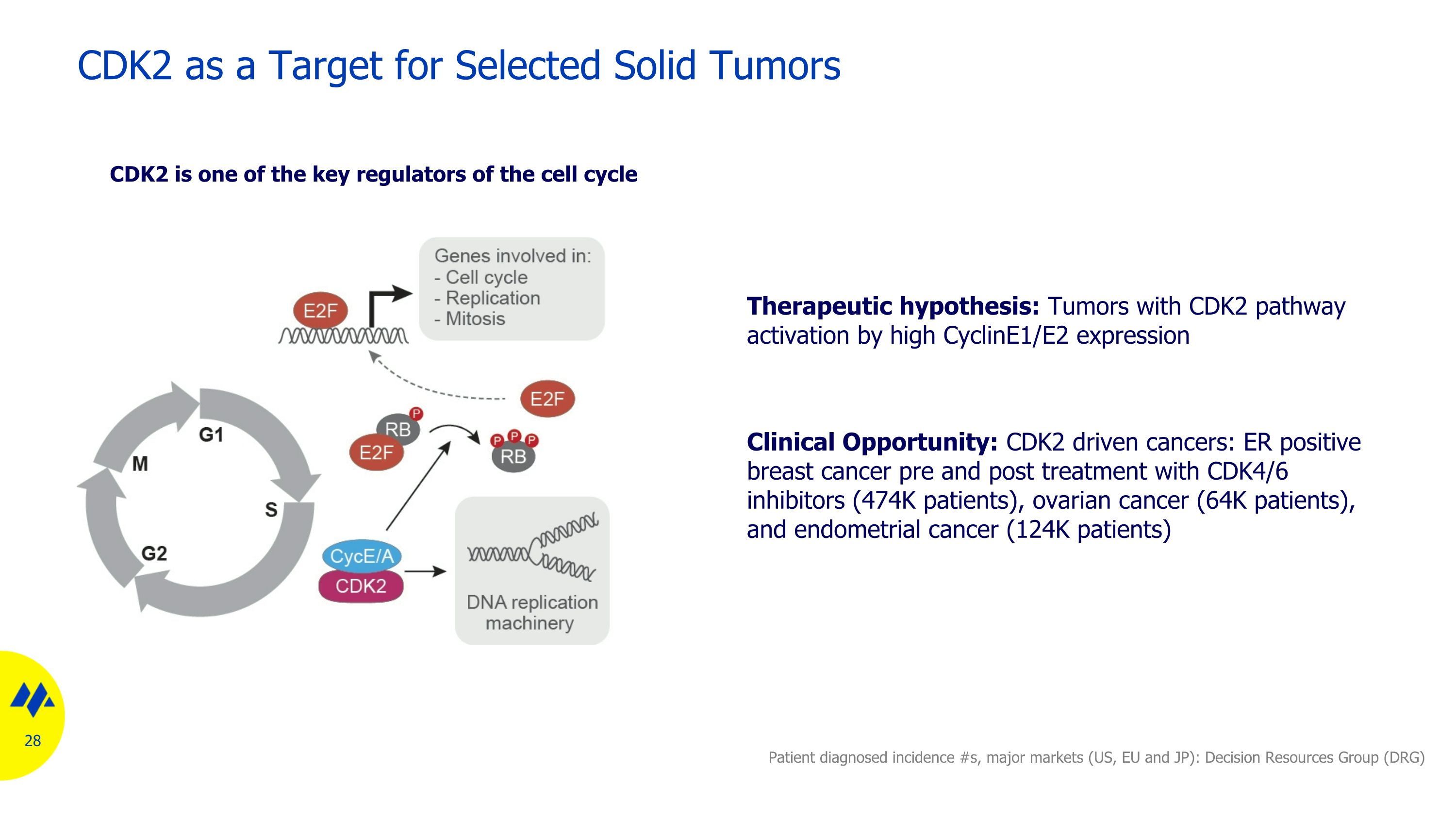
CDK2 as a Target for Selected Solid Tumors Therapeutic hypothesis: Tumors with CDK2 pathway activation by high CyclinE1/E2 expression Clinical Opportunity: CDK2 driven cancers: ER positive breast cancer pre and post treatment with CDK4/6 inhibitors (474K patients), ovarian cancer (64K patients), and endometrial cancer (124K patients) CDK2 is one of the key regulators of the cell cycle Patient diagnosed incidence #s, major markets (US, EU and JP): Decision Resources Group (DRG)
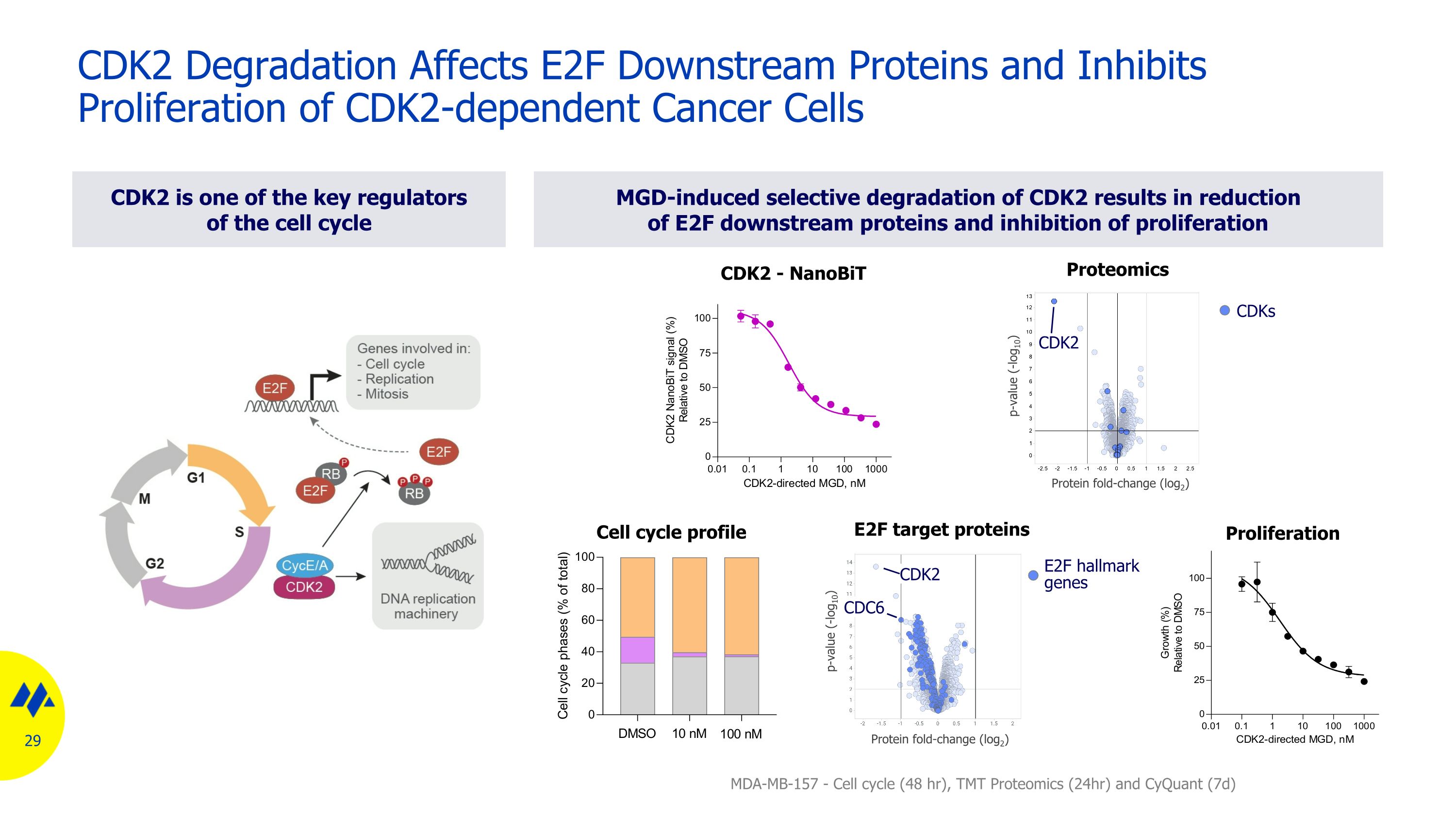
CDK2 Degradation Affects E2F Downstream Proteins and Inhibits Proliferation of CDK2-dependent Cancer Cells p-value (-log10) Protein fold-change (log2) p-value (-log10) Protein fold-change (log2) E2F hallmark genes CDK2 CDC6 MGD-induced selective degradation of CDK2 results in reduction of E2F downstream proteins and inhibition of proliferation CDK2 is one of the key regulators of the cell cycle Proliferation MDA-MB-157 - Cell cycle (48 hr), TMT Proteomics (24hr) and CyQuant (7d) CDK2 - NanoBiT E2F target proteins CDKs CDK2 Proteomics Cell cycle profile
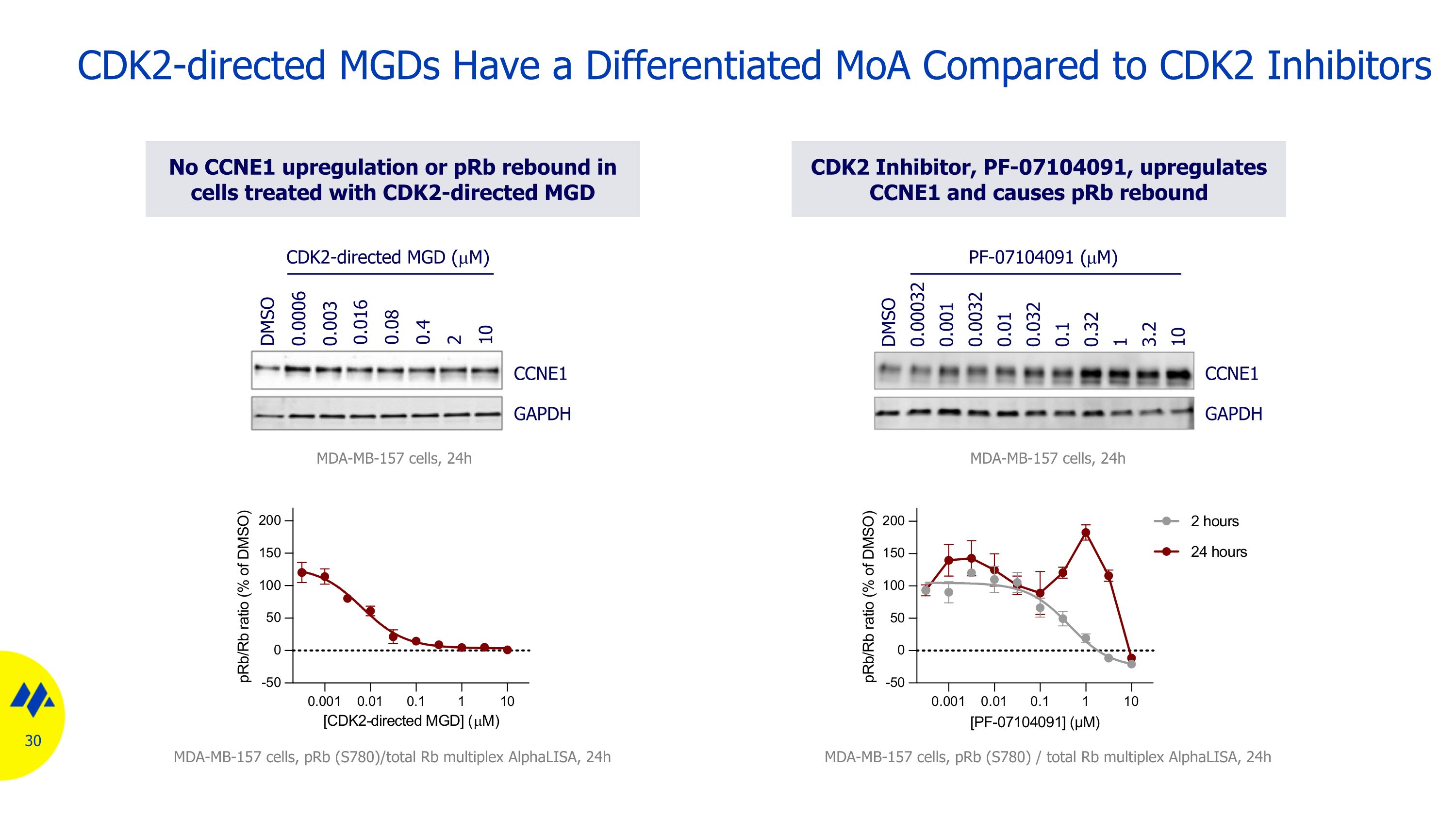
CDK2-directed MGDs Have a Differentiated MoA Compared to CDK2 Inhibitors CDK2 Inhibitor, PF-07104091, upregulates CCNE1 and causes pRb rebound MDA-MB-157 cells, pRb (S780) / total Rb multiplex AlphaLISA, 24h 0.032 1 3.2 10 DMSO 0.1 0.32 0.01 0.0032 0.001 0.00032 CCNE1 GAPDH PF-07104091 (mM) No CCNE1 upregulation or pRb rebound in cells treated with CDK2-directed MGD MDA-MB-157 cells, pRb (S780)/total Rb multiplex AlphaLISA, 24h CCNE1 GAPDH 0.003 0.4 2 10 DMSO 0.016 0.08 0.0006 MDA-MB-157 cells, 24h CDK2-directed MGD (mM) MDA-MB-157 cells, 24h

Inflammation and Immunology (I&I) Programs
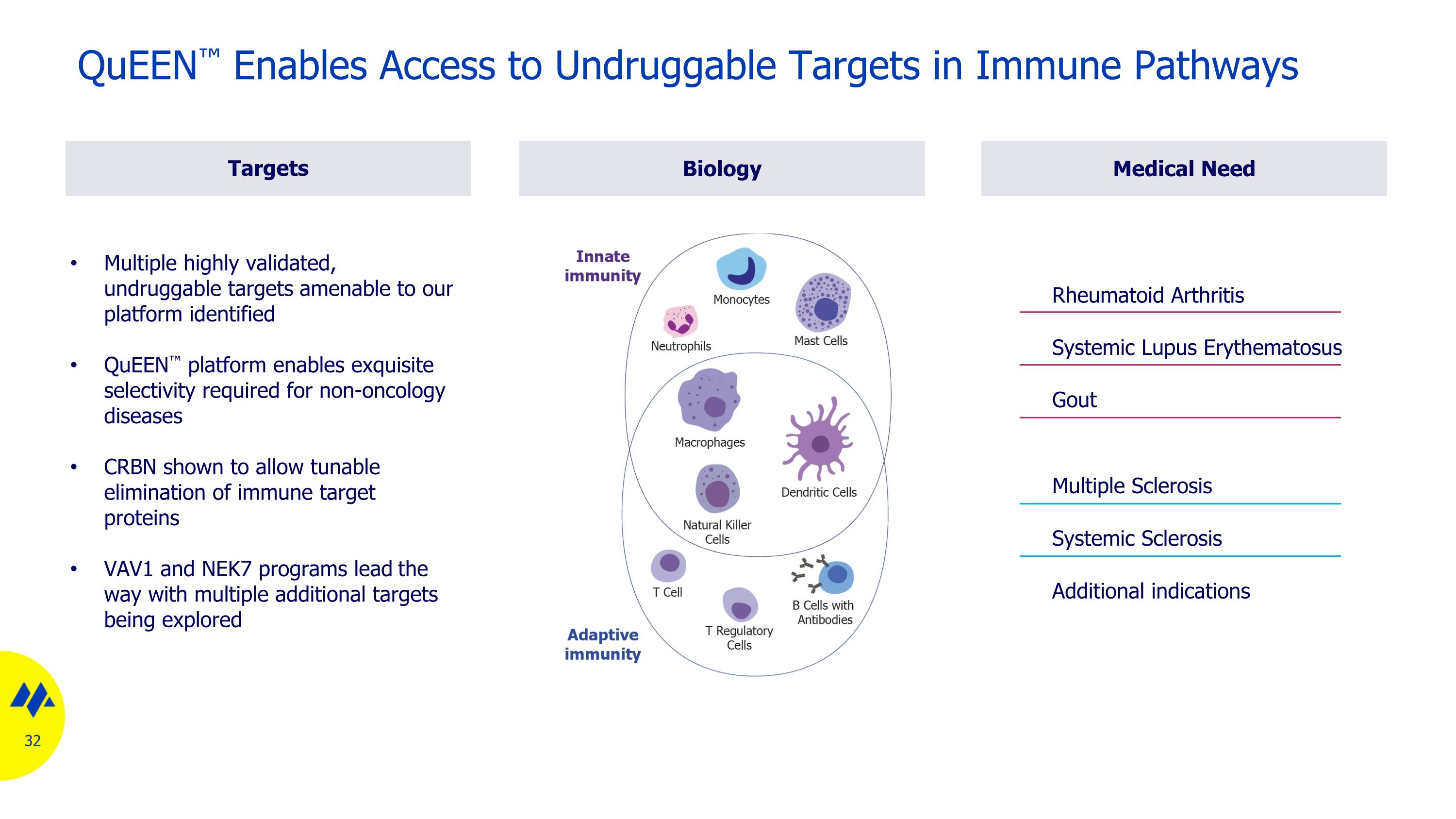
Biology Multiple highly validated, undruggable targets amenable to our platform identified QuEEN™ platform enables exquisite selectivity required for non-oncology diseases CRBN shown to allow tunable elimination of immune target proteins VAV1 and NEK7 programs lead the way with multiple additional targets being explored QuEEN™ Enables Access to Undruggable Targets in Immune Pathways Medical Need Targets Additional indications Systemic Sclerosis Multiple Sclerosis Rheumatoid Arthritis Systemic Lupus Erythematosus Gout

VAV1 Program
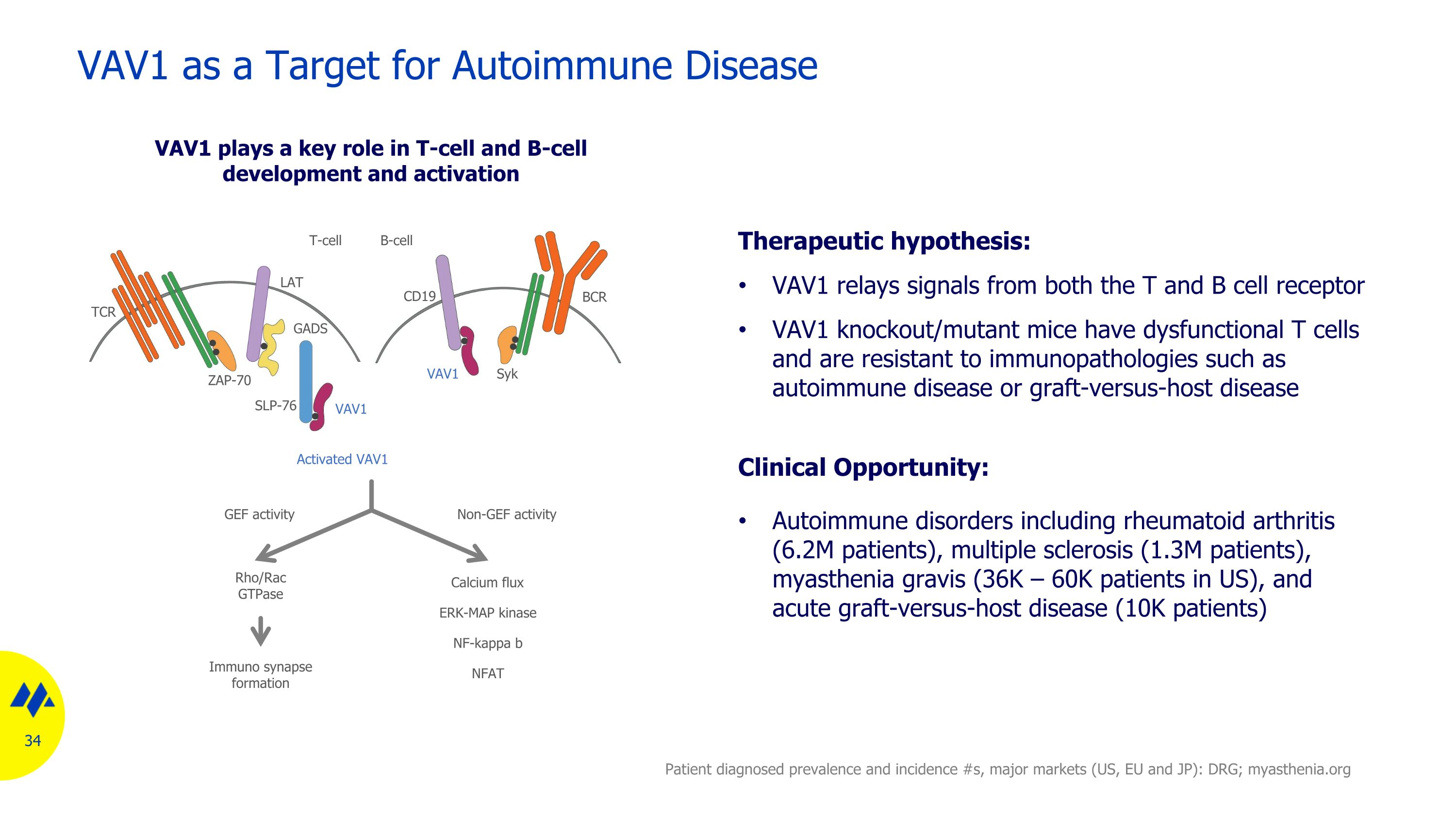
VAV1 as a Target for Autoimmune Disease Therapeutic hypothesis: VAV1 relays signals from both the T and B cell receptor VAV1 knockout/mutant mice have dysfunctional T cells and are resistant to immunopathologies such as autoimmune disease or graft-versus-host disease Clinical Opportunity: Autoimmune disorders including rheumatoid arthritis (6.2M patients), multiple sclerosis (1.3M patients), myasthenia gravis (36K – 60K patients in US), and acute graft-versus-host disease (10K patients) VAV1 plays a key role in T-cell and B-cell development and activation Activated VAV1 B-cell T-cell BCR TCR VAV1 VAV1 Rho/Rac GTPase Calcium flux ERK-MAP kinase NF-kappa b NFAT GEF activity Non-GEF activity Immuno synapse formation CD19 LAT ZAP-70 SLP-76 Syk GADS Patient diagnosed prevalence and incidence #s, major markets (US, EU and JP): DRG; myasthenia.org
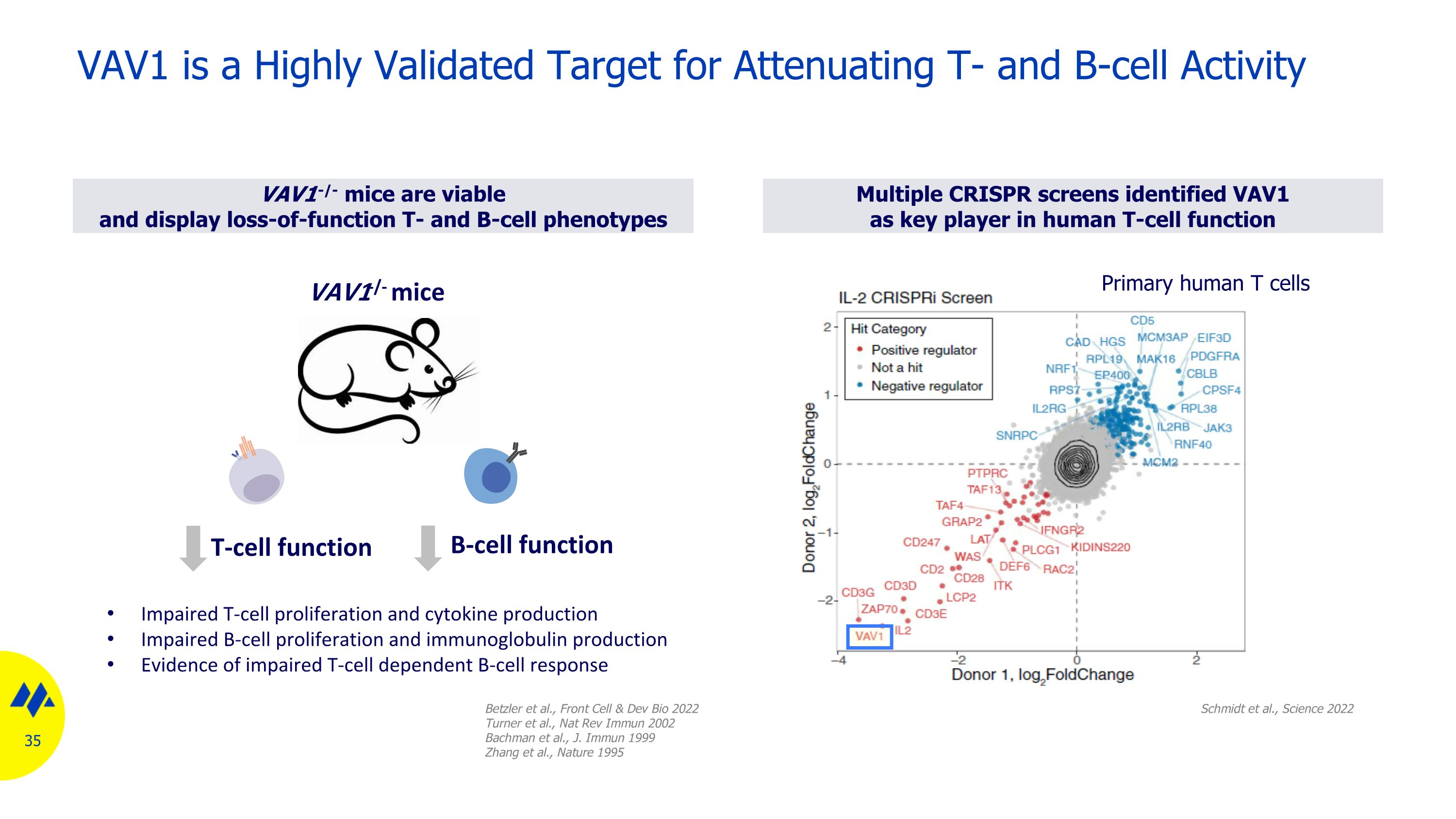
VAV1-/- mice are viable and display loss-of-function T- and B-cell phenotypes Multiple CRISPR screens identified VAV1 as key player in human T-cell function VAV1 is a Highly Validated Target for Attenuating T- and B-cell Activity Schmidt et al., Science 2022 Primary human T cells Betzler et al., Front Cell & Dev Bio 2022 Turner et al., Nat Rev Immun 2002 Bachman et al., J. Immun 1999 Zhang et al., Nature 1995 T-cell function B-cell function VAV1-/- mice Impaired T-cell proliferation and cytokine production Impaired B-cell proliferation and immunoglobulin production Evidence of impaired T-cell dependent B-cell response
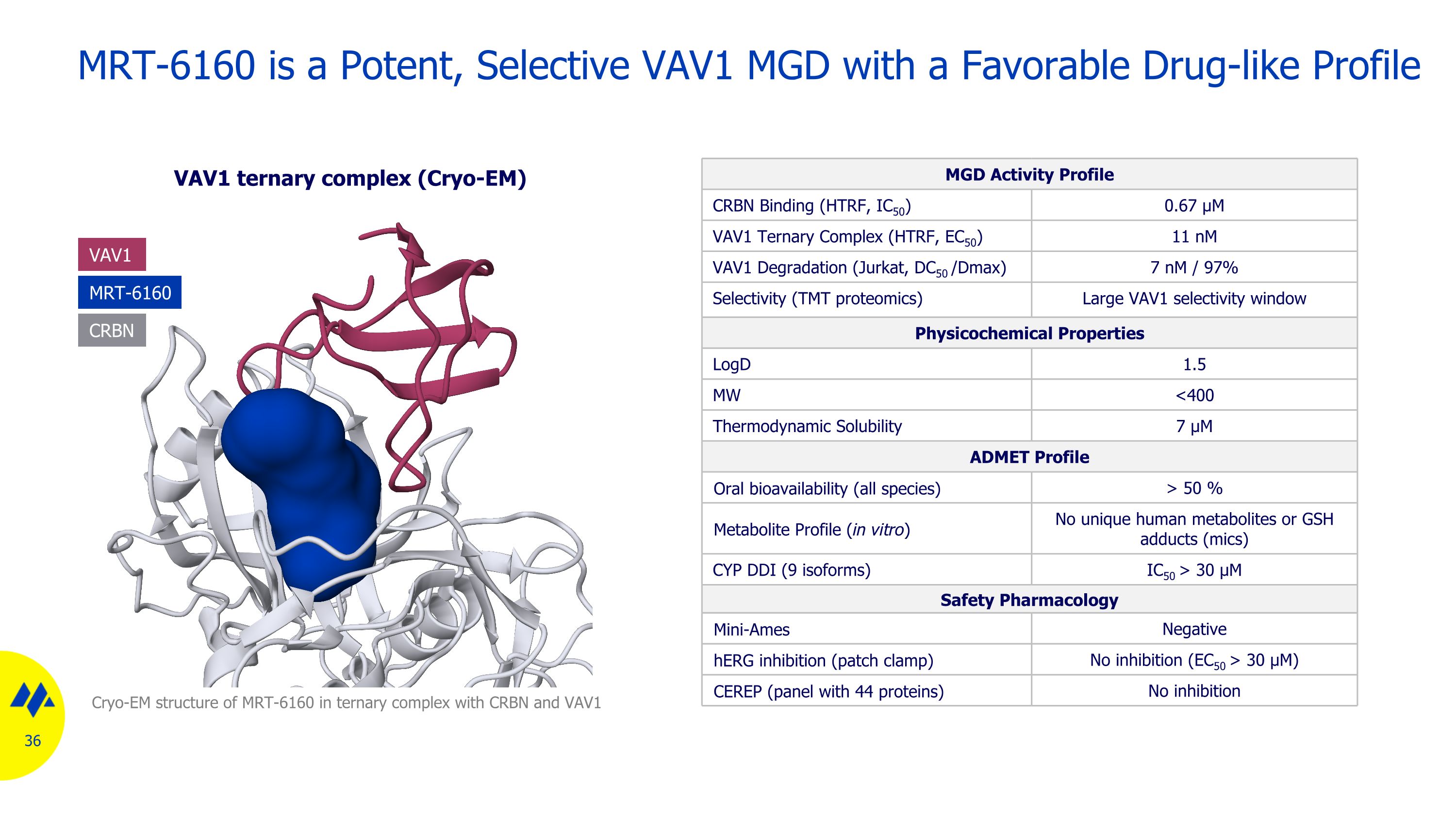
MRT-6160 is a Potent, Selective VAV1 MGD with a Favorable Drug-like Profile VAV1 ternary complex (Cryo-EM) MGD Activity Profile CRBN Binding (HTRF, IC50) 0.67 µM VAV1 Ternary Complex (HTRF, EC50) 11 nM VAV1 Degradation (Jurkat, DC50 /Dmax) 7 nM / 97% Selectivity (TMT proteomics) Large VAV1 selectivity window Physicochemical Properties LogD 1.5 MW <400 Thermodynamic Solubility 7 µM ADMET Profile Oral bioavailability (all species) > 50 % Metabolite Profile (in vitro) No unique human metabolites or GSH adducts (mics) CYP DDI (9 isoforms) IC50 > 30 μM Safety Pharmacology Mini-Ames Negative hERG inhibition (patch clamp) No inhibition (EC50 > 30 µM) CEREP (panel with 44 proteins) No inhibition Cryo-EM structure of MRT-6160 in ternary complex with CRBN and VAV1 MRT-6160 VAV1 CRBN
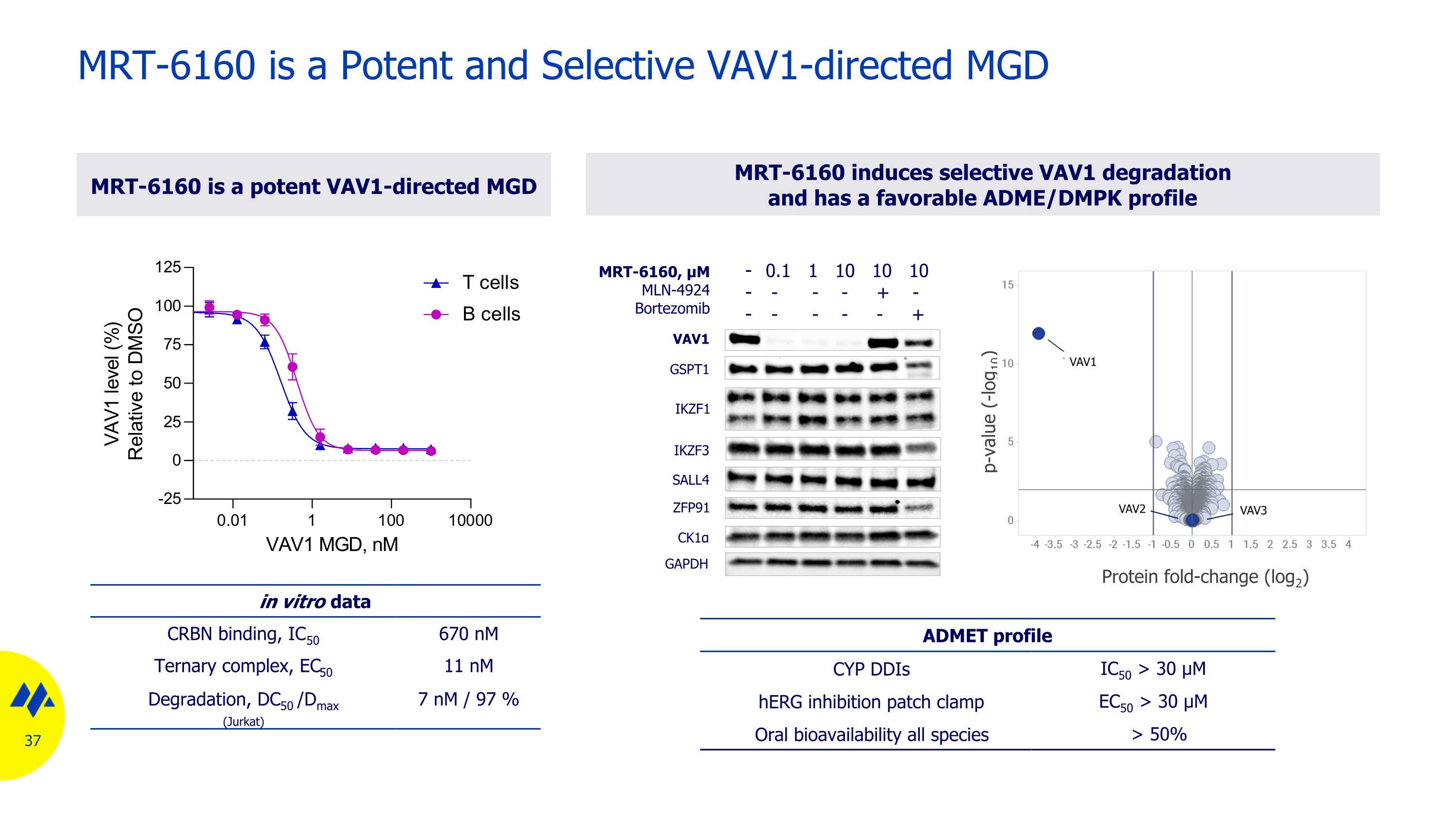
MRT-6160 is a Potent and Selective VAV1-directed MGD in vitro data CRBN binding, IC50 670 nM Ternary complex, EC50 11 nM Degradation, DC50 /Dmax (Jurkat) 7 nM / 97 % MRT-6160 induces selective VAV1 degradation and has a favorable ADME/DMPK profile MRT-6160 is a potent VAV1-directed MGD ADMET profile CYP DDIs IC50 > 30 µM hERG inhibition patch clamp EC50 > 30 µM Oral bioavailability all species > 50% p-value (-log10) Protein fold-change (log2) GAPDH CK1α ZFP91 GSPT1 IKZF1 VAV1 SALL4 IKZF3 MRT-6160, μM MLN-4924 Bortezomib 0.1 1 10 10 10 - - - + - - - - - +
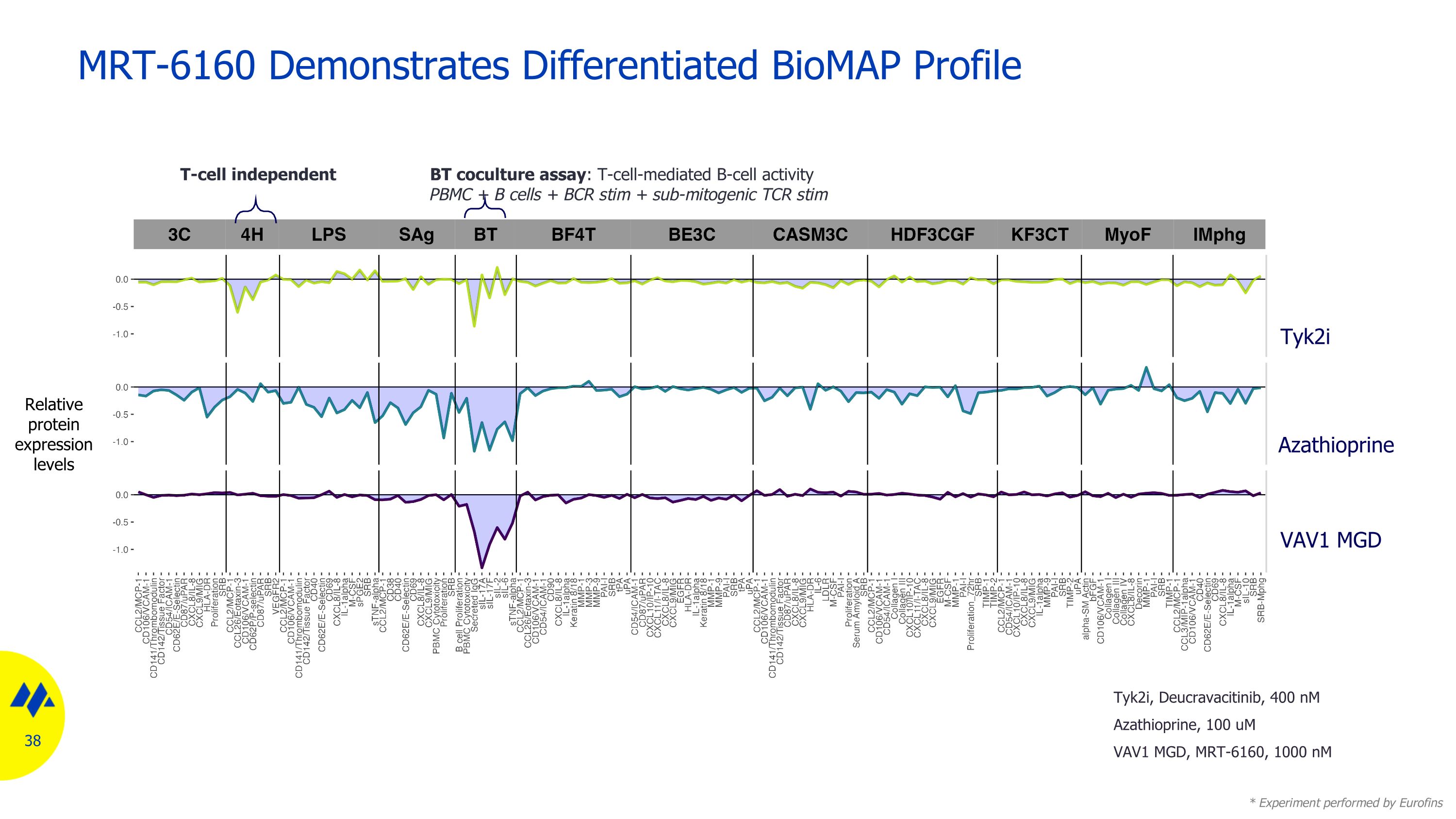
MRT-6160 Demonstrates Differentiated BioMAP Profile Relative protein expression levels Azathioprine VAV1 MGD Tyk2i BT coculture assay: T-cell-mediated B-cell activity PBMC + B cells + BCR stim + sub-mitogenic TCR stim * Experiment performed by Eurofins T-cell independent Tyk2i, Deucravacitinib, 400 nM Azathioprine, 100 uM VAV1 MGD, MRT-6160, 1000 nM
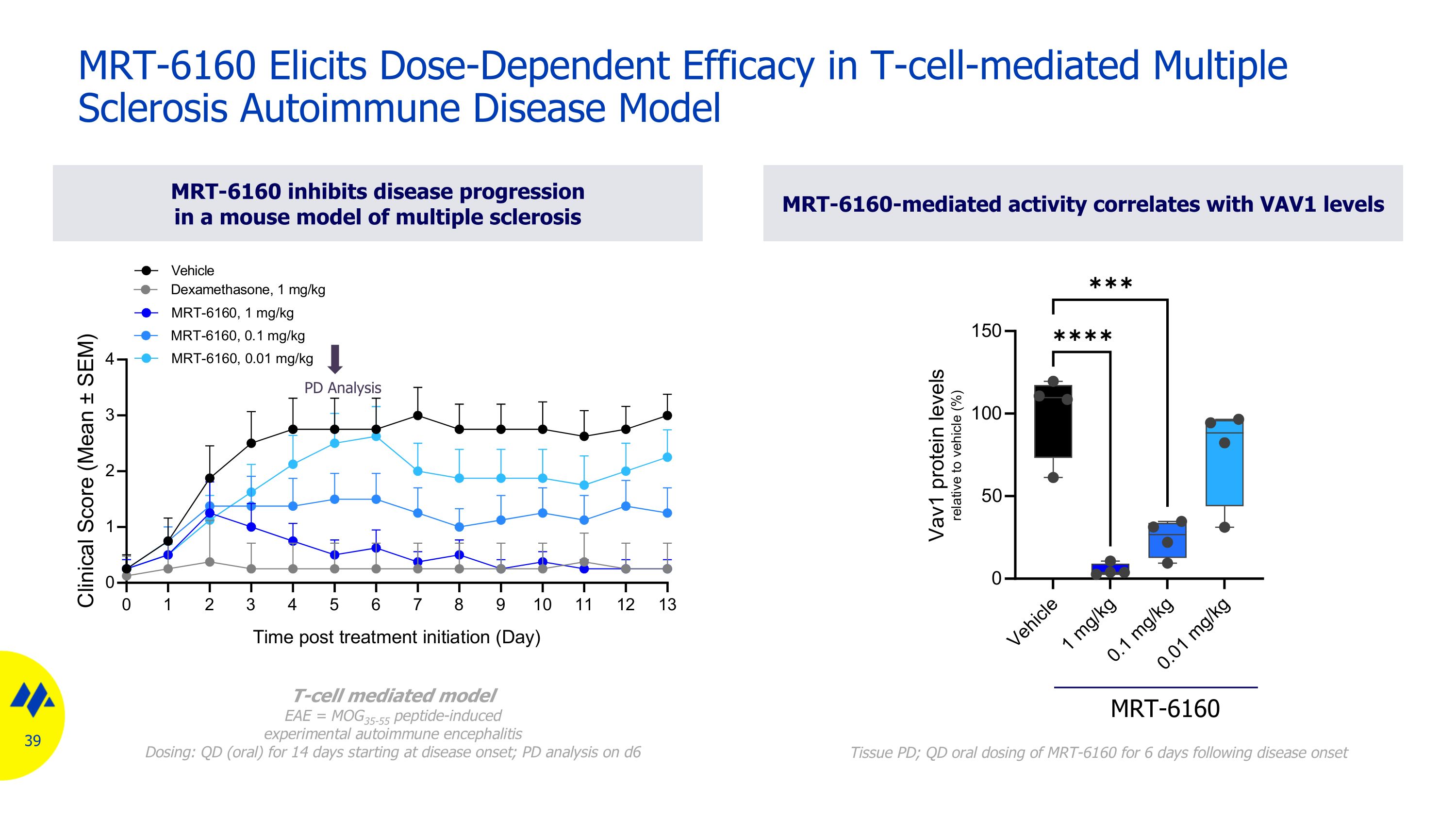
PD Analysis MRT-6160 inhibits disease progression in a mouse model of multiple sclerosis MRT-6160-mediated activity correlates with VAV1 levels MRT-6160 Elicits Dose-Dependent Efficacy in T-cell-mediated Multiple Sclerosis Autoimmune Disease Model T-cell mediated model EAE = MOG35-55 peptide-induced experimental autoimmune encephalitis Dosing: QD (oral) for 14 days starting at disease onset; PD analysis on d6 MRT-6160 Tissue PD; QD oral dosing of MRT-6160 for 6 days following disease onset
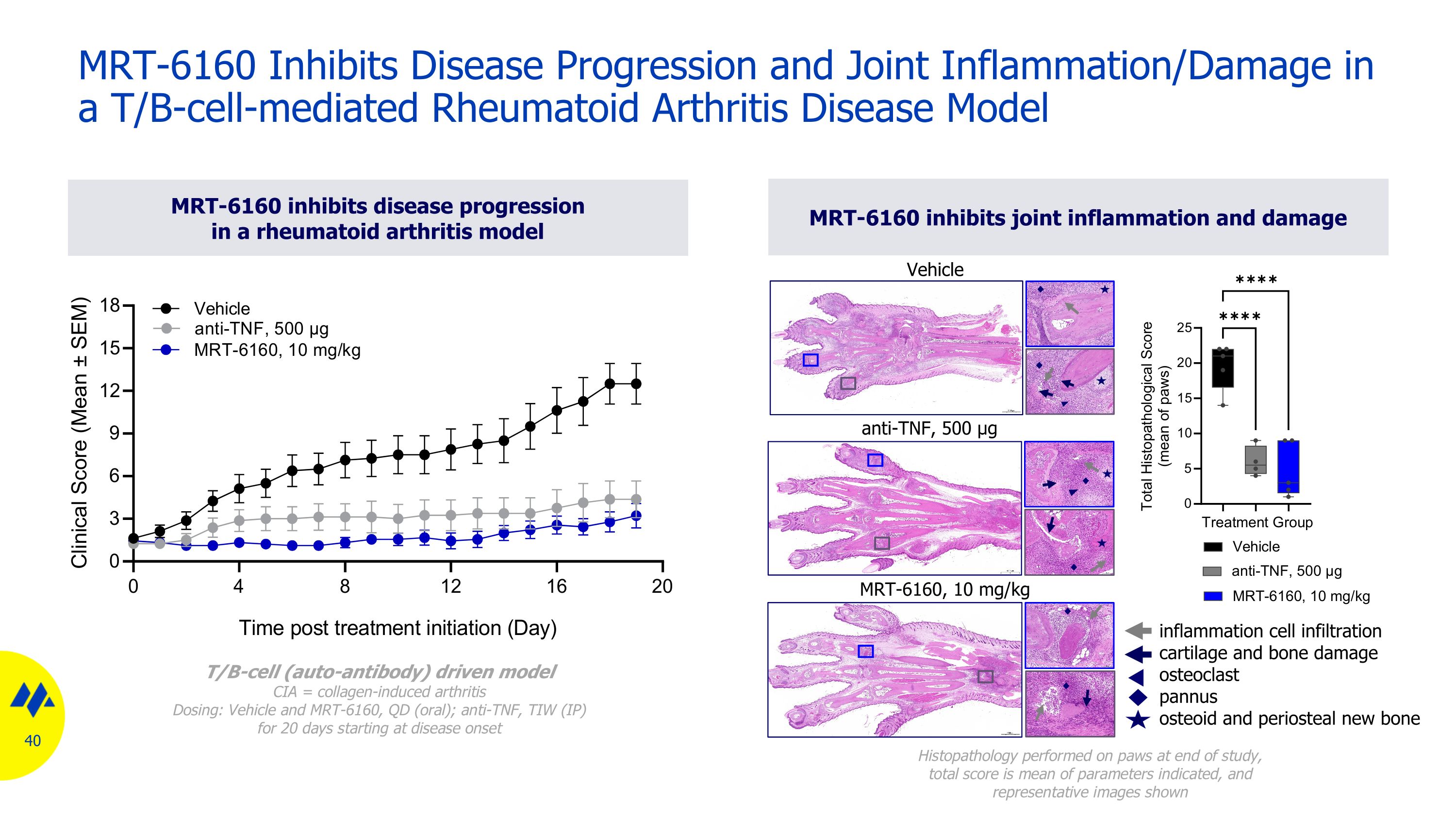
MRT-6160 inhibits disease progression in a rheumatoid arthritis model MRT-6160 inhibits joint inflammation and damage MRT-6160 Inhibits Disease Progression and Joint Inflammation/Damage in a T/B-cell-mediated Rheumatoid Arthritis Disease Model T/B-cell (auto-antibody) driven model CIA = collagen-induced arthritis Dosing: Vehicle and MRT-6160, QD (oral); anti-TNF, TIW (IP) for 20 days starting at disease onset Vehicle anti-TNF, 500 µg MRT-6160, 10 mg/kg Histopathology performed on paws at end of study, total score is mean of parameters indicated, and representative images shown inflammation cell infiltration cartilage and bone damage osteoclast pannus osteoid and periosteal new bone
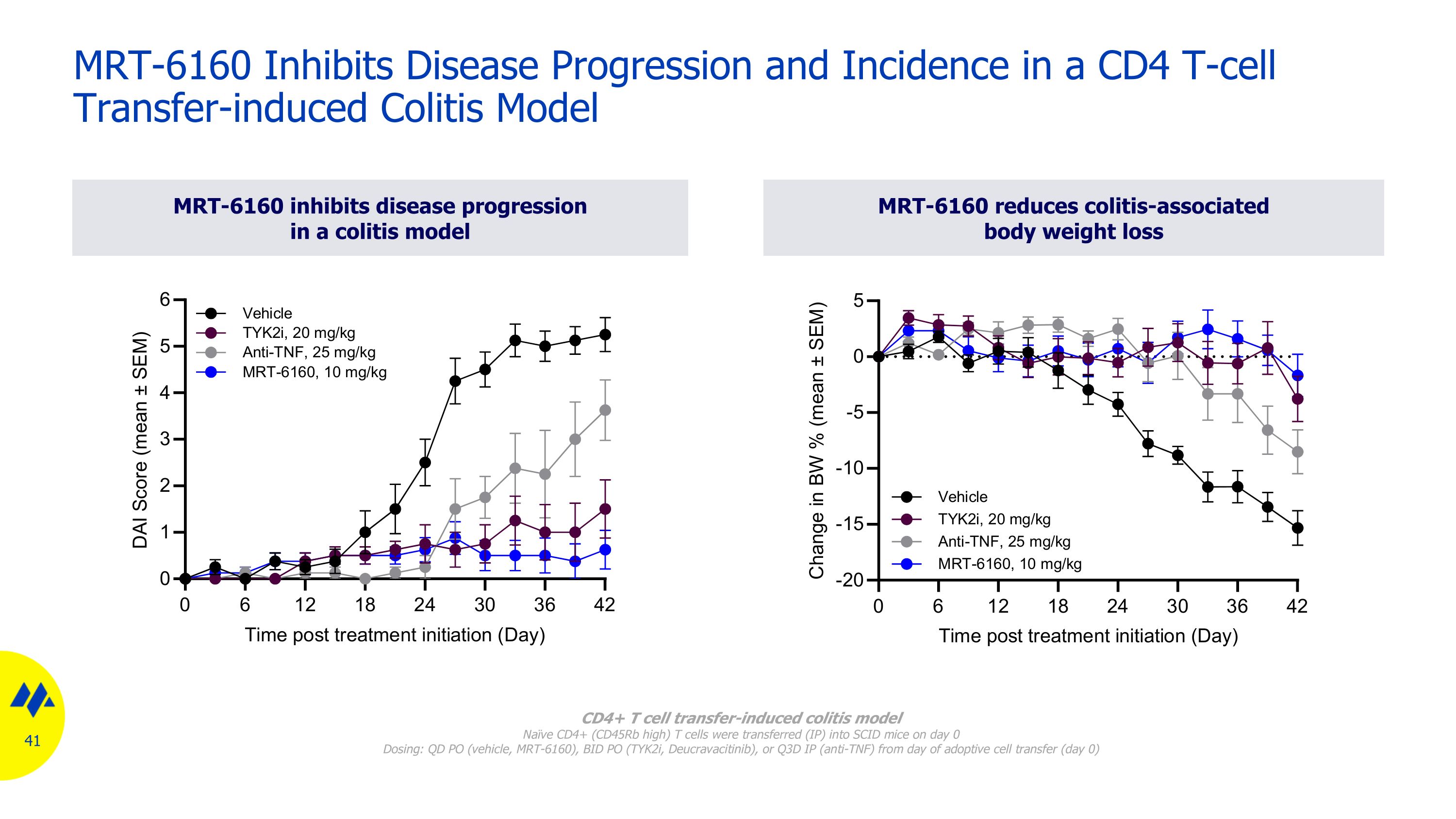
MRT-6160 inhibits disease progression in a colitis model MRT-6160 reduces colitis-associated body weight loss MRT-6160 Inhibits Disease Progression and Incidence in a CD4 T-cell Transfer-induced Colitis Model CD4+ T cell transfer-induced colitis model Naïve CD4+ (CD45Rb high) T cells were transferred (IP) into SCID mice on day 0 Dosing: QD PO (vehicle, MRT-6160), BID PO (TYK2i, Deucravacitinib), or Q3D IP (anti-TNF) from day of adoptive cell transfer (day 0)
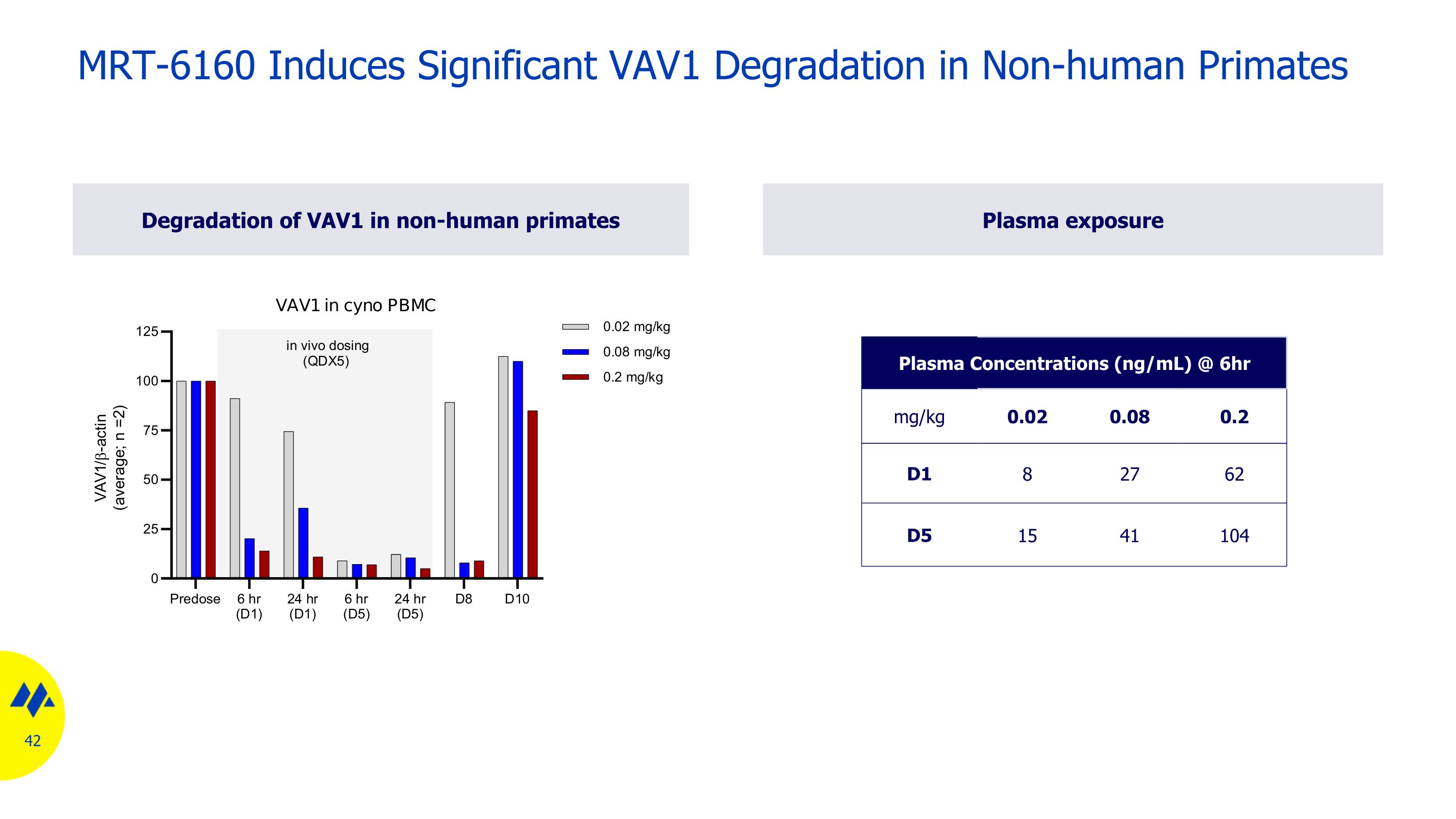
Degradation of VAV1 in non-human primates Plasma exposure MRT-6160 Induces Significant VAV1 Degradation in Non-human Primates Plasma Concentrations (ng/mL) @ 6hr mg/kg 0.02 0.08 0.2 D1 8 27 62 D5 15 41 104

MRT-6160 – Our VAV1-directed MGD for Autoimmune Disease MRT-6160 is a highly selective VAV1-directed MGD designed through our QuEENTM platform and the first MGD being developed specifically for a non-oncology indication Attenuates multiple aspects of T- and B-cell receptor signaling in relevant preclinical models Engages VAV1 through a novel binding mode and non-canonical degron In vivo inhibition of disease progression shown in EAE, CIA and IBD mouse models IND filing expected in 1H 2024 Current clinical plan developed with the goal of providing early insights into safety, PK and PD, and proof-of-concept (POC) regarding differentiated effects on key immunomodulatory signaling pathways Potential to address significant unmet opportunities in multiple autoimmune disorders including dermatology, IBD, multiple sclerosis and rheumatology

NEK7 Program
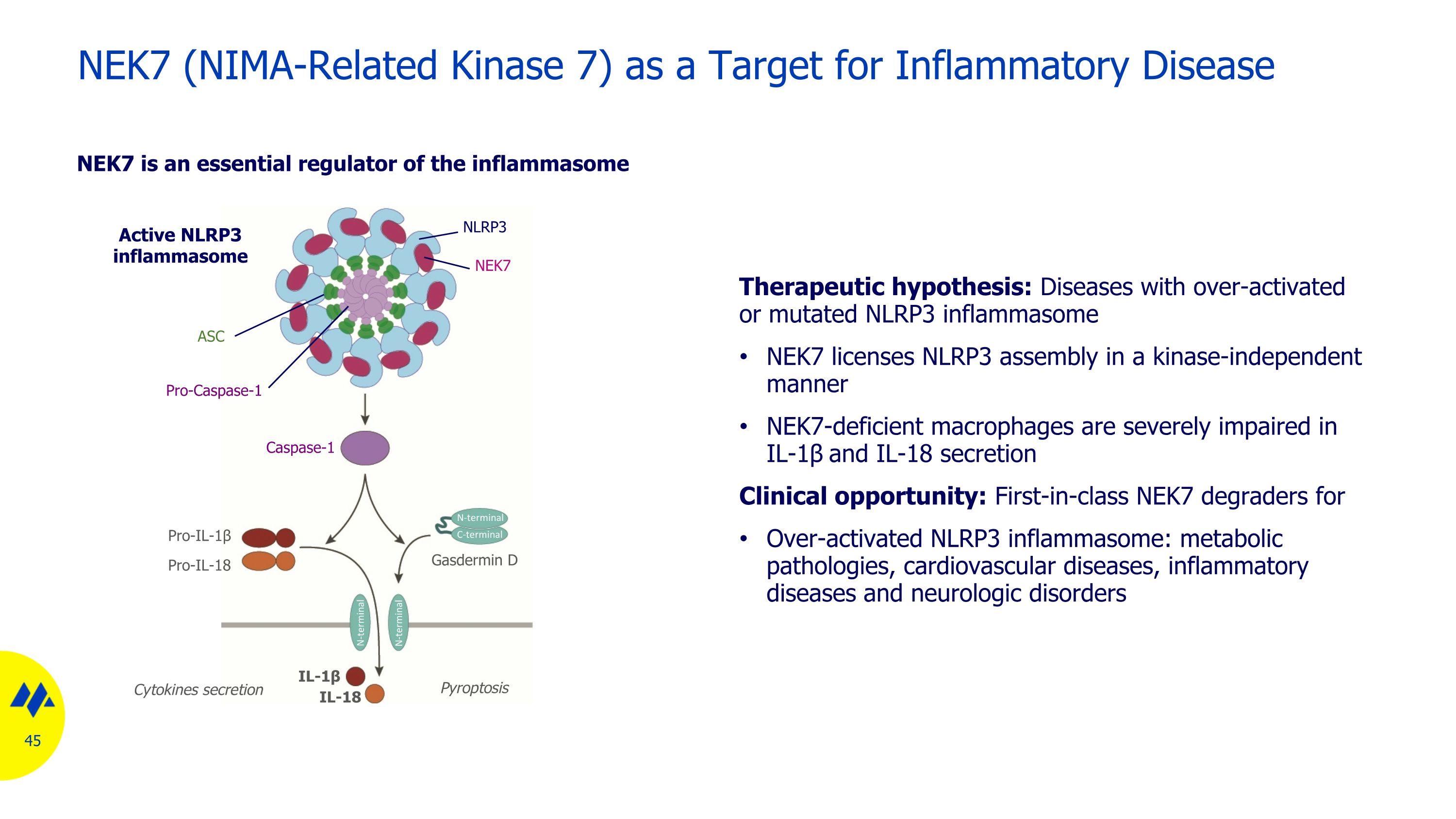
NEK7 (NIMA-Related Kinase 7) as a Target for Inflammatory Disease NEK7 is an essential regulator of the inflammasome Pyroptosis Gasdermin D Active NLRP3 inflammasome Cytokines secretion Pro-IL-1β Pro-IL-18 IL-18 IL-1β NEK7 NLRP3 Pro-Caspase-1 ASC N-terminal C-terminal N-terminal Caspase-1 N-terminal Therapeutic hypothesis: Diseases with over-activated or mutated NLRP3 inflammasome NEK7 licenses NLRP3 assembly in a kinase-independent manner NEK7-deficient macrophages are severely impaired in IL-1β and IL-18 secretion Clinical opportunity: First-in-class NEK7 degraders for Over-activated NLRP3 inflammasome: metabolic pathologies, cardiovascular diseases, inflammatory diseases and neurologic disorders
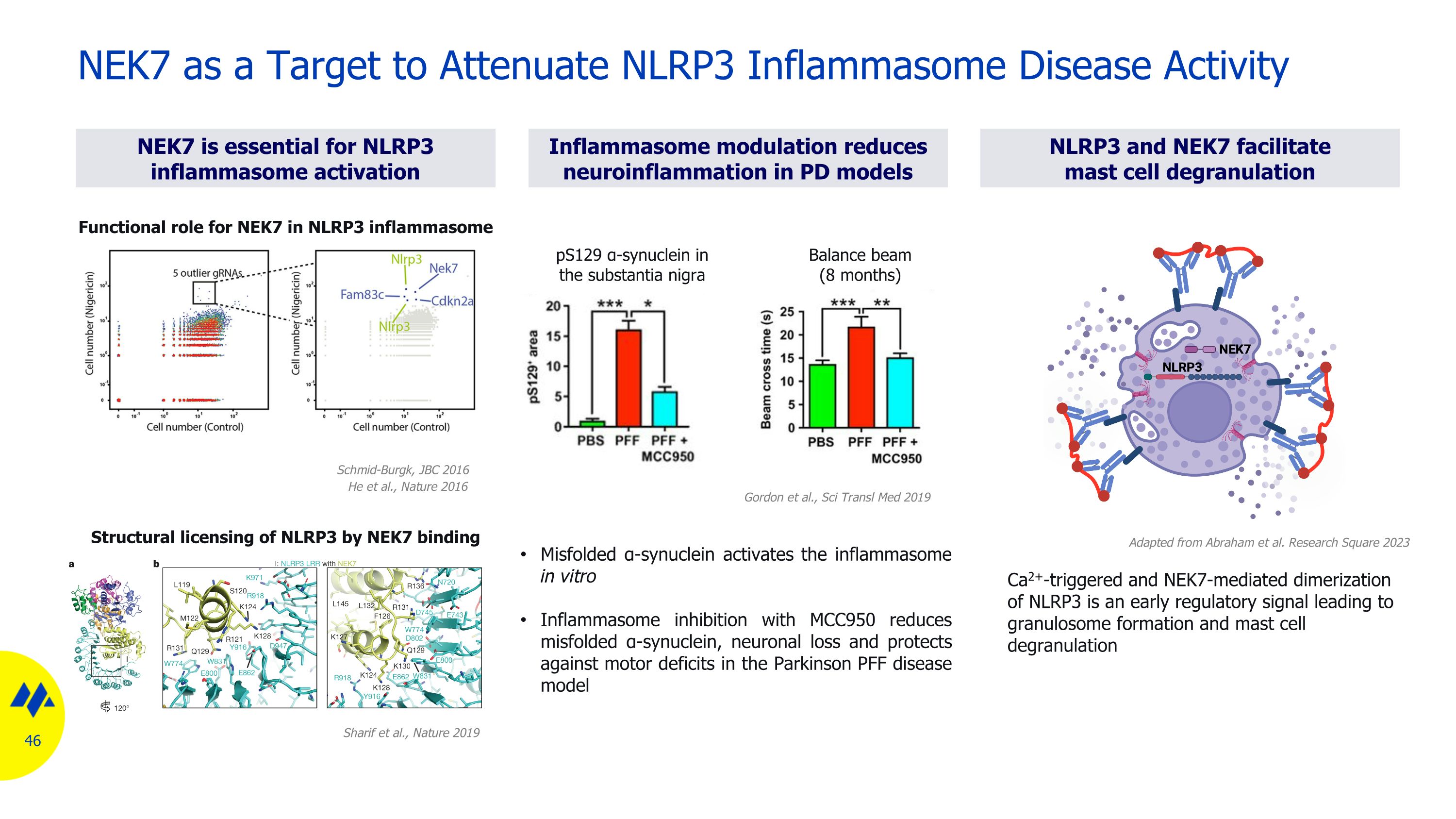
NEK7 as a Target to Attenuate NLRP3 Inflammasome Disease Activity Misfolded α-synuclein activates the inflammasome in vitro Inflammasome inhibition with MCC950 reduces misfolded α-synuclein, neuronal loss and protects against motor deficits in the Parkinson PFF disease model pS129 α-synuclein in the substantia nigra Balance beam (8 months) Ca2+-triggered and NEK7-mediated dimerization of NLRP3 is an early regulatory signal leading to granulosome formation and mast cell degranulation Inflammasome modulation reduces neuroinflammation in PD models NEK7 is essential for NLRP3 inflammasome activation NLRP3 and NEK7 facilitate mast cell degranulation Sharif et al., Nature 2019 Structural licensing of NLRP3 by NEK7 binding Schmid-Burgk, JBC 2016 He et al., Nature 2016 Functional role for NEK7 in NLRP3 inflammasome Gordon et al., Sci Transl Med 2019 Adapted from Abraham et al. Research Square 2023
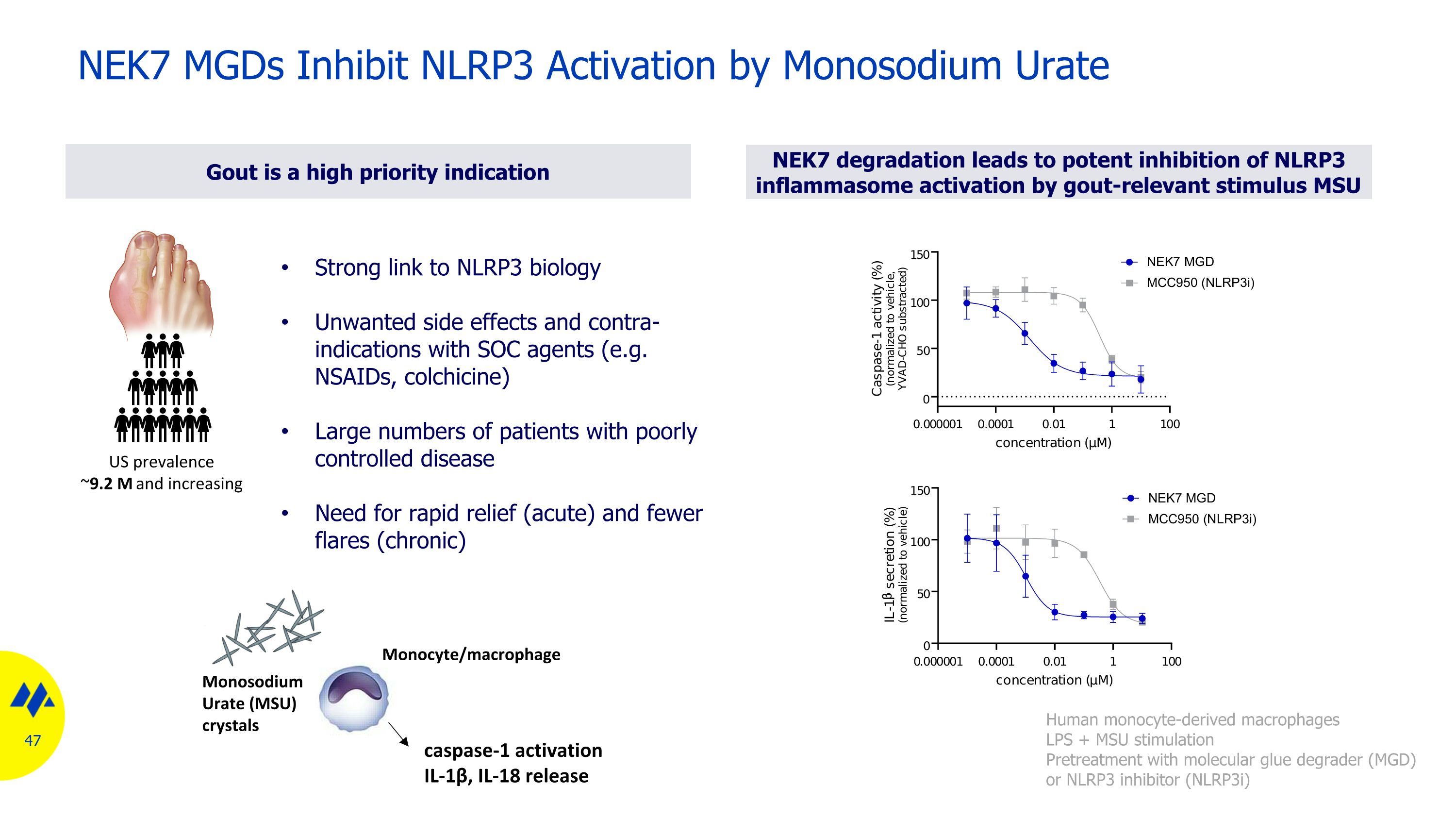
NEK7 MGDs Inhibit NLRP3 Activation by Monosodium Urate US prevalence ~9.2 M and increasing Monocyte/macrophage caspase-1 activation IL-1β, IL-18 release Monosodium�Urate (MSU) crystals Strong link to NLRP3 biology Unwanted side effects and contra-indications with SOC agents (e.g. NSAIDs, colchicine) Large numbers of patients with poorly controlled disease Need for rapid relief (acute) and fewer flares (chronic) Gout is a high priority indication Human monocyte-derived macrophages LPS + MSU stimulation Pretreatment with molecular glue degrader (MGD) or NLRP3 inhibitor (NLRP3i) NEK7 degradation leads to potent inhibition of NLRP3 inflammasome activation by gout-relevant stimulus MSU

MRT-8102 is a NEK7-Directed MGD With Favorable Drug-like Properties MGD Activity Profile CRBN Binding (HTRF, IC50) 0.2 µM NEK7 Degradation (CAL51, DC50 /Dmax) 10 nM / 89% Selectivity (TMT proteomics) Excellent selectivity profile in different cell lines Physicochemical Properties LogD 1.47 MW <450 Thermodynamic Solubility 166 µM ADMET Profile Oral Bioavailability Yes Metabolite Profile (in vitro) No unique human metabolites or GSH adducts (mics) Safety Pharmacology Mini-Ames Negative hERG (patch clamp) No inhibition (EC50> 30 µM) CEREP (panel with 44 proteins) No inhibition NEK7 Ternary Complex (Crystal Structure) MRT-8102 NEK7 CRBN
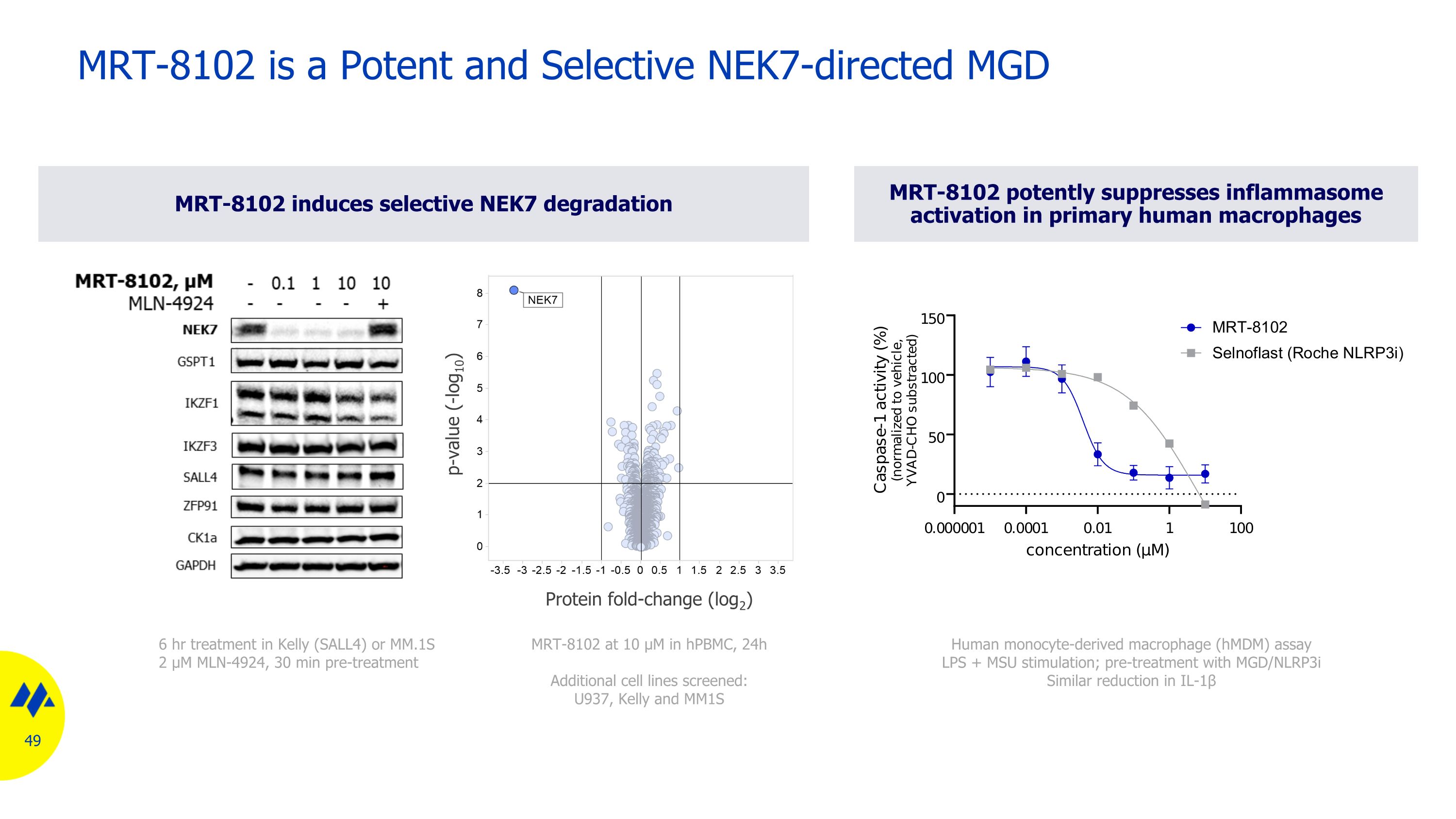
MRT-8102 potently suppresses inflammasome activation in primary human macrophages MRT-8102 induces selective NEK7 degradation p-value (-log10) Protein fold-change (log2) MRT-8102 at 10 µM in hPBMC, 24h Additional cell lines screened: U937, Kelly and MM1S 6 hr treatment in Kelly (SALL4) or MM.1S 2 µM MLN-4924, 30 min pre-treatment Human monocyte-derived macrophage (hMDM) assay LPS + MSU stimulation; pre-treatment with MGD/NLRP3i Similar reduction in IL-1β MRT-8102 is a Potent and Selective NEK7-directed MGD

MRT-8102 inhibits caspase-1 activity in hMDMs after stimulation Human monocyte-derived macrophage (hMDM) assay Pre-incubation with MGD/NLRP3i; stimulated with LPS + MSU Supernatants analyzed post-stimulation MRT-8102 Potently Inhibits NLRP3 Inflammasome-mediated Activation in Human Monocyte-derived Macrophages MRT-8102 inhibits IL-1β secretion by hMDMs after stimulation MRT-8102 inhibits IL-18 secretion by hMDMs after stimulation
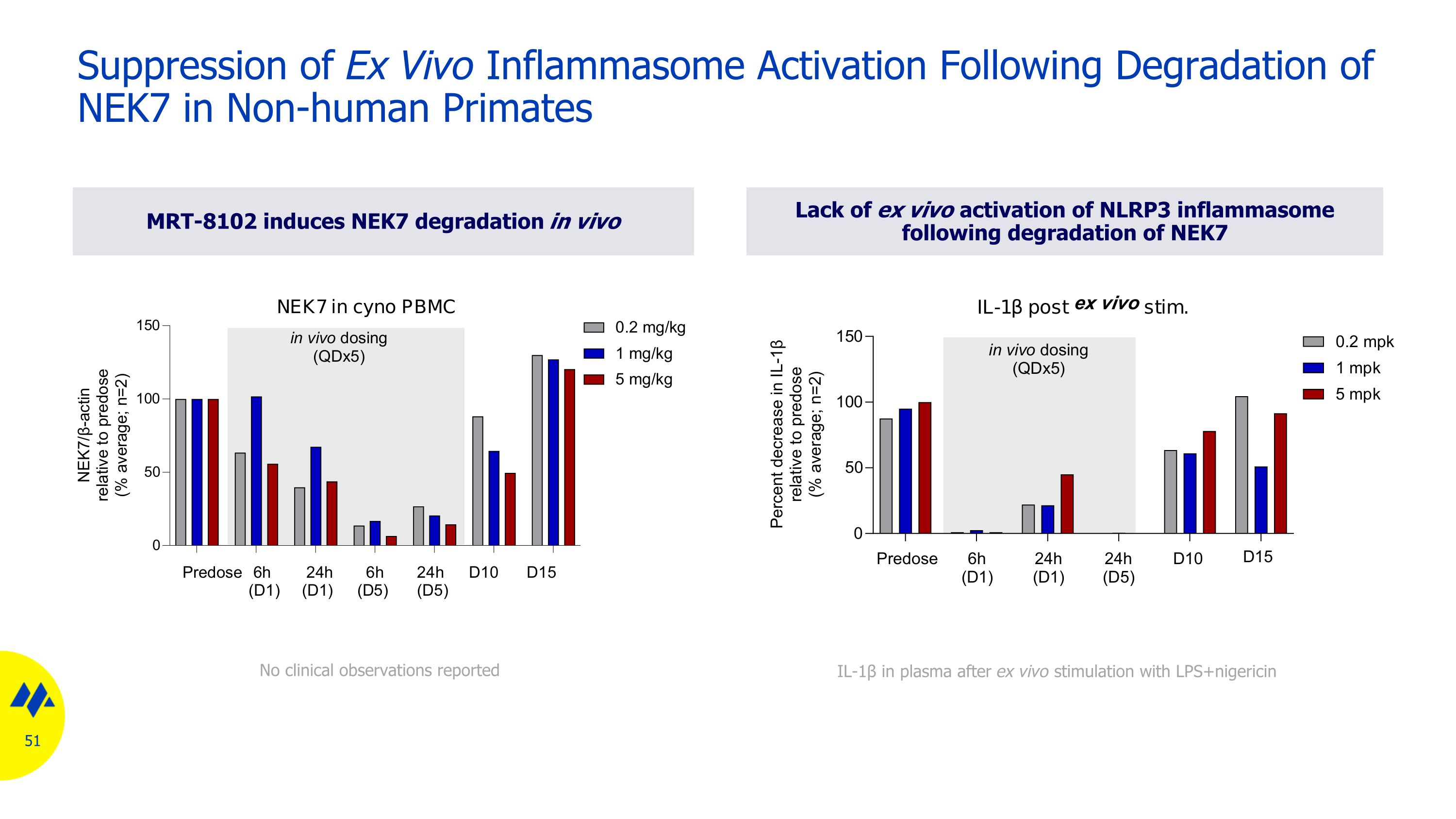
MRT-8102 induces NEK7 degradation in vivo Lack of ex vivo activation of NLRP3 inflammasome following degradation of NEK7 Suppression of Ex Vivo Inflammasome Activation Following Degradation of NEK7 in Non-human Primates No clinical observations reported IL-1β in plasma after ex vivo stimulation with LPS+nigericin

Sickle Cell Disease Program
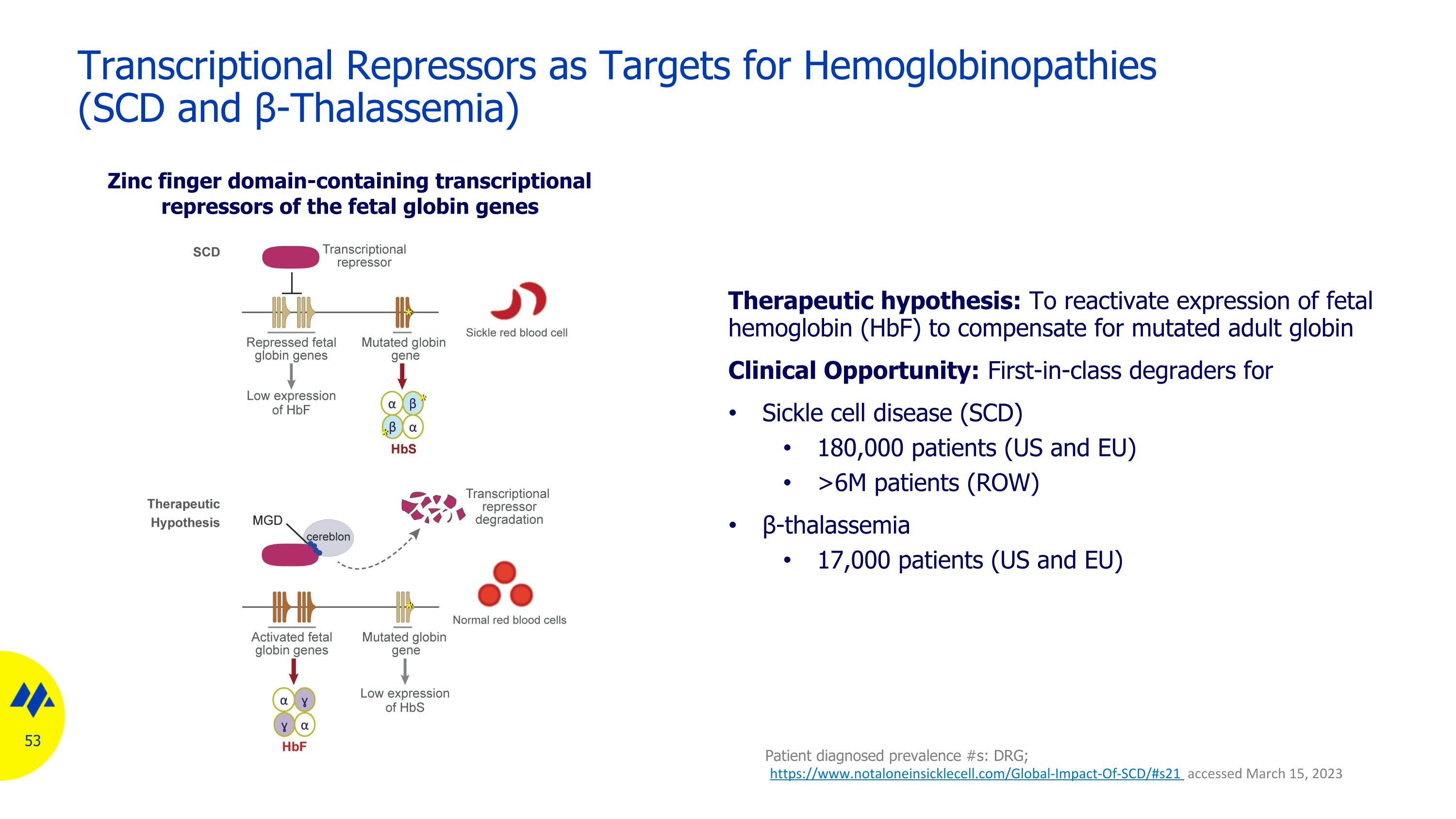
Transcriptional Repressors as Targets for Hemoglobinopathies �(SCD and β-Thalassemia) Therapeutic hypothesis: To reactivate expression of fetal hemoglobin (HbF) to compensate for mutated adult globin Clinical Opportunity: First-in-class degraders for Sickle cell disease (SCD) 180,000 patients (US and EU) >6M patients (ROW) β-thalassemia 17,000 patients (US and EU) Zinc finger domain-containing transcriptional repressors of the fetal globin genes Patient diagnosed prevalence #s: DRG; https://www.notaloneinsicklecell.com/Global-Impact-Of-SCD/#s21 accessed March 15, 2023

Our MGD Library A rationally designed library as a starting point to tackle unprecedented neosubstrates
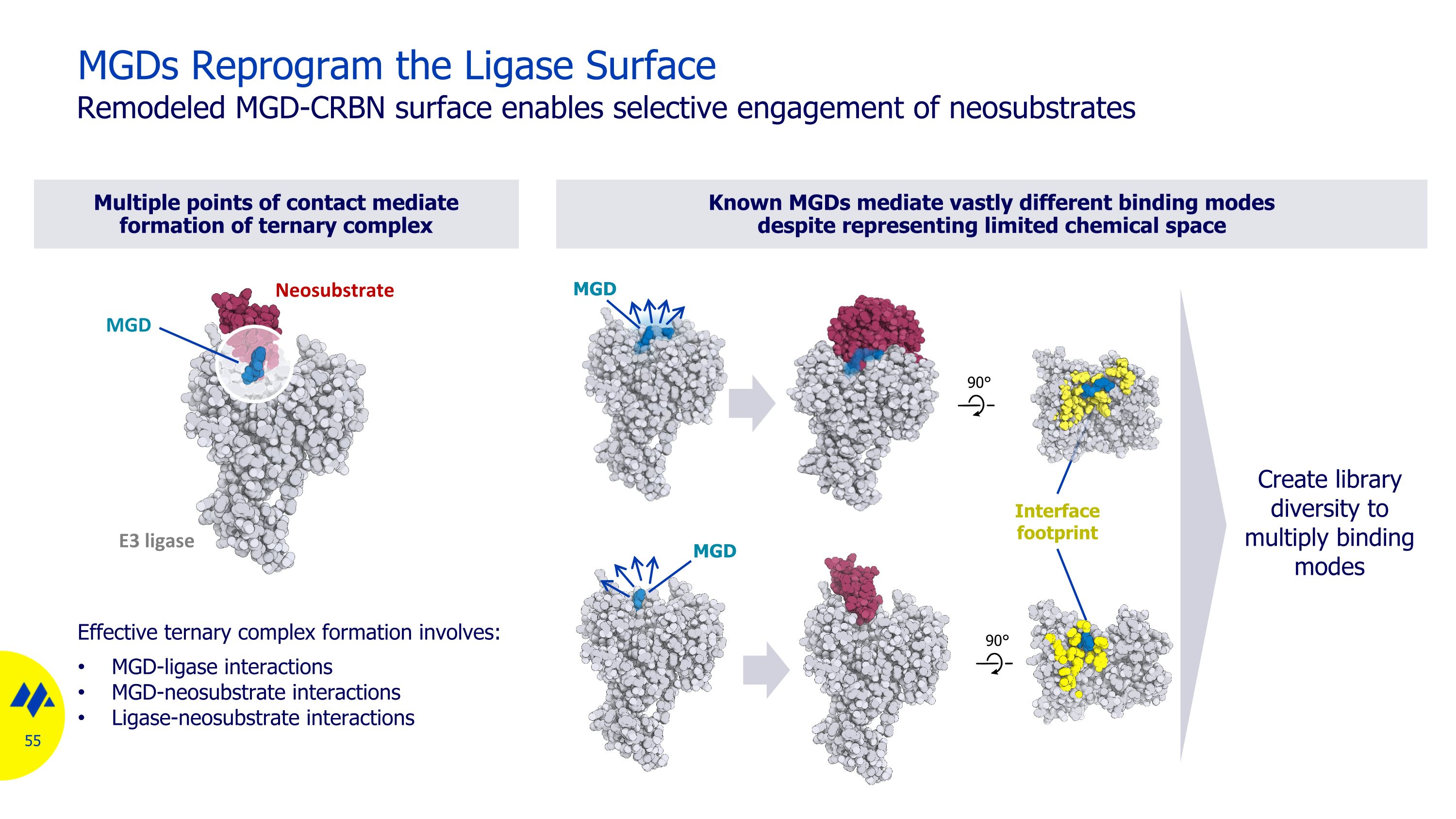
Multiple points of contact mediate formation of ternary complex Known MGDs mediate vastly different binding modes despite representing limited chemical space MGDs Reprogram the Ligase Surface Remodeled MGD-CRBN surface enables selective engagement of neosubstrates Effective ternary complex formation involves: MGD-ligase interactions MGD-neosubstrate interactions Ligase-neosubstrate interactions MGD MGD Create library diversity to multiply binding modes Interface footprint E3 ligase Neosubstrate MGD 90° 90°
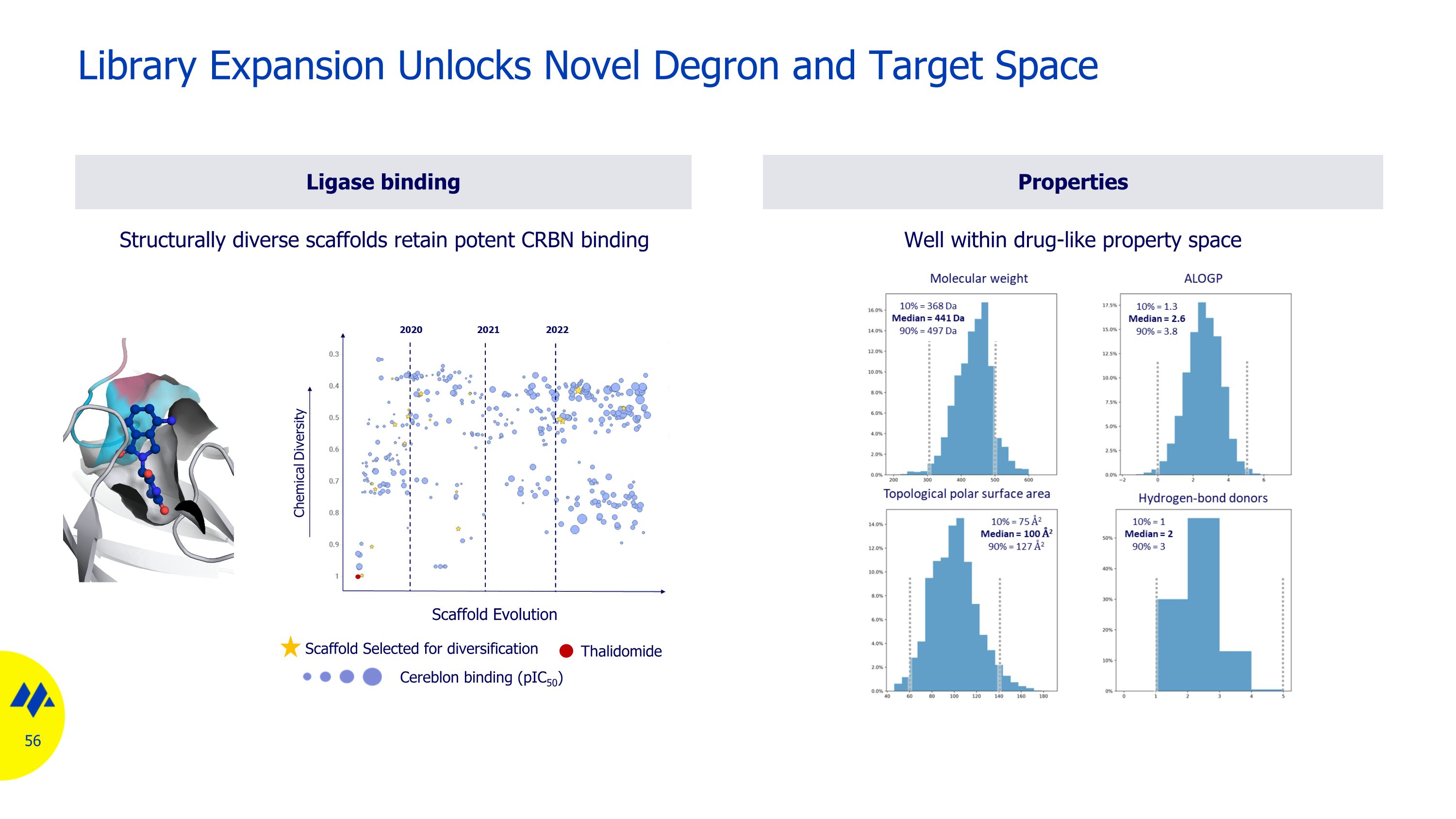
Ligase binding Structurally diverse scaffolds retain potent CRBN binding Library Expansion Unlocks Novel Degron and Target Space Scaffold Evolution Thalidomide Scaffold Selected for diversification Cereblon binding (pIC50) Properties Well within drug-like property space
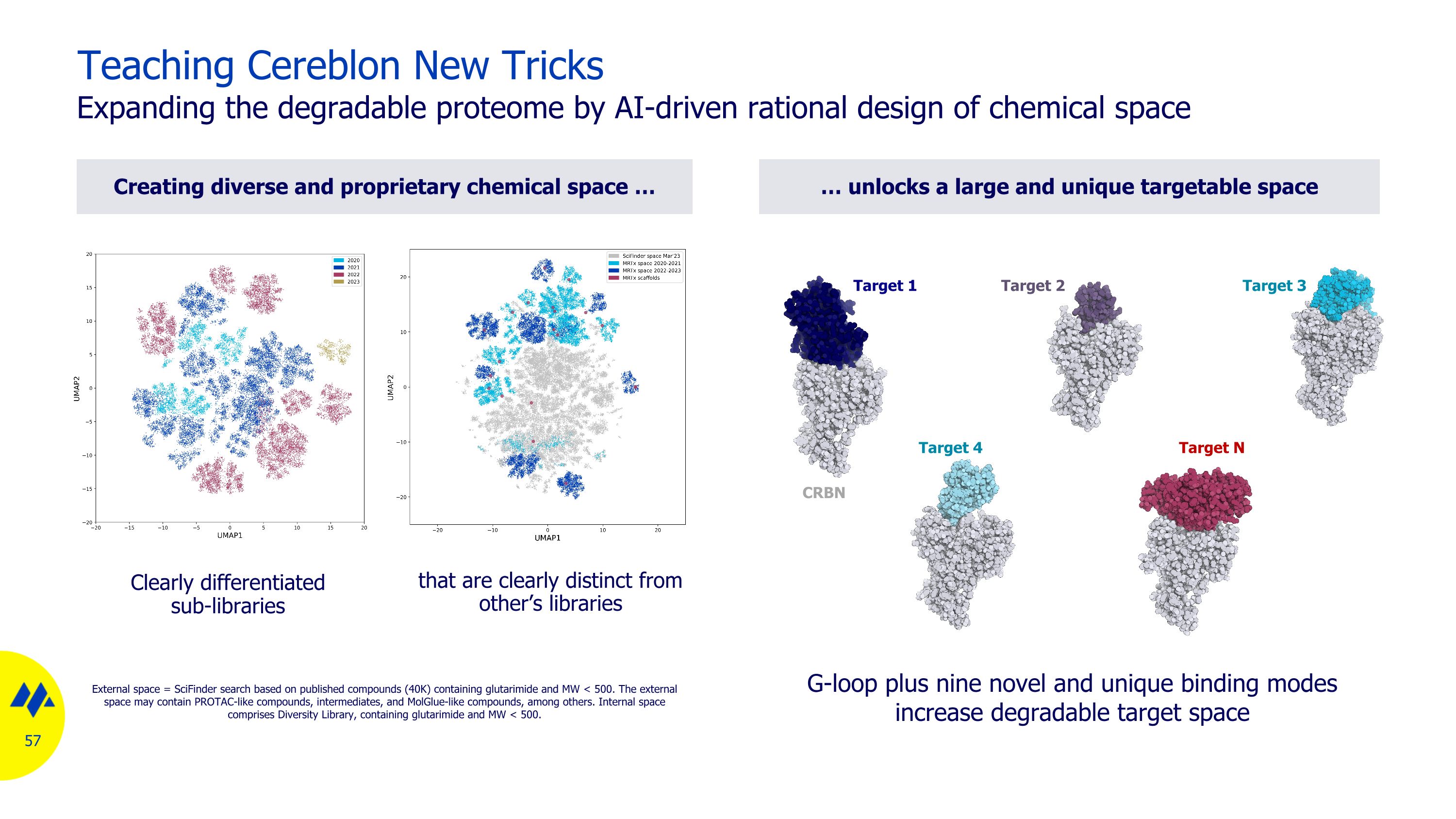

Creating diverse and proprietary chemical space … … unlocks a large and unique targetable space Clearly differentiated sub-libraries that are clearly distinct from other’s libraries External space = SciFinder search based on published compounds (40K) containing glutarimide and MW < 500. The external space may contain PROTAC-like compounds, intermediates, and MolGlue-like compounds, among others. Internal space comprises Diversity Library, containing glutarimide and MW < 500. Teaching Cereblon New Tricks Expanding the degradable proteome by AI-driven rational design of chemical space Target 1 Target 3 Target 2 Target 4 Target N CRBN G-loop plus nine novel and unique binding modes increase degradable target space

Team

World-Class Leadership Deep expertise in molecular glue discovery, drug development and precision medicine Jullian Jones, �Ph.D., J.D., MBA Chief Business Officer Filip Janku, M.D., Ph.D. Chief Medical Officer Markus Warmuth, M.D. Chief Executive Officer John Castle, Ph.D. Chief Data Scientist & Information Officer Sharon Townson, Ph.D. Chief Technology Officer Owen Wallace, Ph.D. President of Research and Preclinical Development Phil Nickson, Ph.D., J.D. General Counsel Jennifer Champoux Chief People & Operations Officer Magnus Walter, Ph.D. SVP, Chemical Sciences and Process Development

Thank You



























































